Crystal structure of Cel7A from Talaromyces emersonii in complex with cellotetraose
Qin, L., Pereira, J.H., McAndrew, R.P., Simmons, B.A., Sapra, R., Adams, P.D., Sale, K.L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cellobiohydrolase 1 catalytic domain | 437 | Rasamsonia emersonii | Mutation(s): 0 Gene Names: cbh1, cbh1A EC: 3.2.1.91 (PDB Primary Data), 3.2.1 (UniProt) | 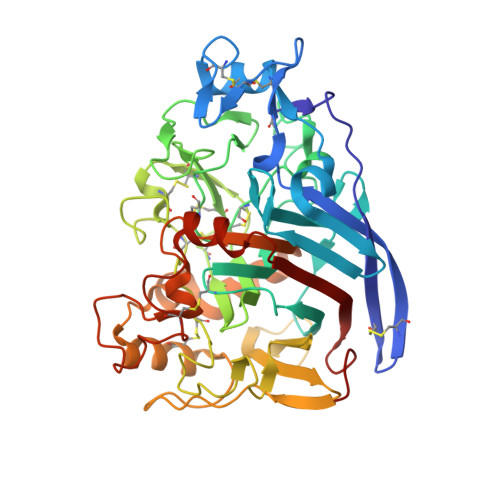 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8TFL9 | ||||
Glycosylation | |||||
| Glycosylation Sites: 2 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | B | 4 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G81315DD GlyCosmos: G81315DD GlyGen: G81315DD | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| beta-D-glucopyranose-(1-4)-beta-D-glucopyranose-(1-4)-beta-D-glucopyranose-(1-4)-beta-D-glucopyranose | C | 4 |  | N/A | |
Glycosylation Resources | |||||
GlyTouCan: G00025MO GlyCosmos: G00025MO GlyGen: G00025MO | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | E [auth A] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | F [auth A], G [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| PCA Query on PCA | A | L-PEPTIDE LINKING | C5 H7 N O3 |  | GLN |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_900011 Query on PRD_900011 | C | beta-cellotetraose | Oligosaccharide / Metabolism |  |
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_900005 Query on PRD_900005 | D | beta-cellobiose | Oligosaccharide / Metabolism |  |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 74.72 | α = 90 |
| b = 74.72 | β = 90 |
| c = 169.922 | γ = 90 |
| Software Name | Purpose |
|---|---|
| BOS | data collection |
| PHASER | phasing |
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |