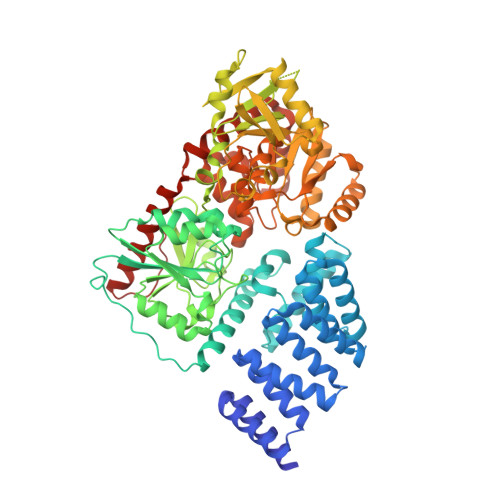
Structure of human O-GlcNAc transferase and its complex with a peptide substrate.
Lazarus, M.B., Nam, Y., Jiang, J., Sliz, P., Walker, S.(2011) Nature 469: 564-567
- PubMed: 21240259 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09638
- Primary Citation Related Structures:
3PE3, 3PE4 - PubMed Abstract:
The essential mammalian enzyme O-linked β-N-acetylglucosamine transferase (O-GlcNAc transferase, here OGT) couples metabolic status to the regulation of a wide variety of cellular signalling pathways by acting as a nutrient sensor. OGT catalyses the transfer of N-acetylglucosamine from UDP-N-acetylglucosamine (UDP-GlcNAc) to serines and threonines of cytoplasmic, nuclear and mitochondrial proteins, including numerous transcription factors, tumour suppressors, kinases, phosphatases and histone-modifying proteins. Aberrant glycosylation by OGT has been linked to insulin resistance, diabetic complications, cancer and neurodegenerative diseases including Alzheimer's. Despite the importance of OGT, the details of how it recognizes and glycosylates its protein substrates are largely unknown. We report here two crystal structures of human OGT, as a binary complex with UDP (2.8 Å resolution) and as a ternary complex with UDP and a peptide substrate (1.95 Å). The structures provide clues to the enzyme mechanism, show how OGT recognizes target peptide sequences, and reveal the fold of the unique domain between the two halves of the catalytic region. This information will accelerate the rational design of biological experiments to investigate OGT's functions; it will also help the design of inhibitors for use as cellular probes and help to assess its potential as a therapeutic target.
- Department of Chemistry and Chemical Biology, Harvard University, Cambridge, Massachusetts 02138, USA.
Organizational Affiliation: