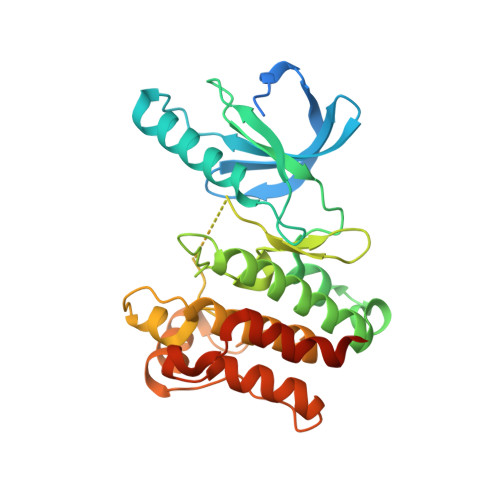
Protein kinase domain of CTR1 from Arabidopsis thaliana promotes ethylene receptor cross talk.
Mayerhofer, H., Panneerselvam, S., Mueller-Dieckmann, J.(2012) J Mol Biology 415: 768-779
- PubMed: 22155294 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.11.046
- Primary Citation Related Structures:
3P86, 3PPZ - PubMed Abstract:
Ethylene controls many aspects of plant growth and development. Signaling by the gaseous phytohormone is initiated by disulfide-linked membrane-bound receptors, and the formation of heteromeric receptor clusters contributes to the broad range of ethylene responsiveness. In Arabidopsis thaliana, the TCS-like ethylene receptors interact with the cytosolic serine/threonine kinase constitutive triple response 1 (CTR1), a proposed mitogen-activated protein kinase kinase kinase. In the absence of the hormone, the receptor and therefore CTR1 are active. Hence, ethylene acts as an inverse agonist of its signaling pathway. The three-dimensional structures of the active, triphosphorylated and the unphosphorylated, inactive kinase domain of CTR1 in complex with staurosporine illustrate the conformational rearrangements that form the basis of activity regulation. Additionally, in analytical ultracentrifugation experiments, active kinase domains form back-to-back dimers, while inactive and activation loop variants are monomers. Together with a front-to-front activation interface, the active protein kinase dimers thereby engage in interactions that promote CTR1-mediated cross talk between ethylene receptor clusters. This model provides a structural foundation for the observed high sensitivity of plants to ethylene.
- EMBL Hamburg Outstation, Hamburg, Germany.
Organizational Affiliation: