Crystal structures of active and inhibitor-bound human Casp6
Mueller, I., Lamers, M.B.A.C., Ritchie, A.J., Dominguez, C., Munoz, I., Maillard, M., Kiselyov, A.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Caspase-6 | 157 | Homo sapiens | Mutation(s): 0 Gene Names: CASP6, MCH2 EC: 3.4.22.59 | 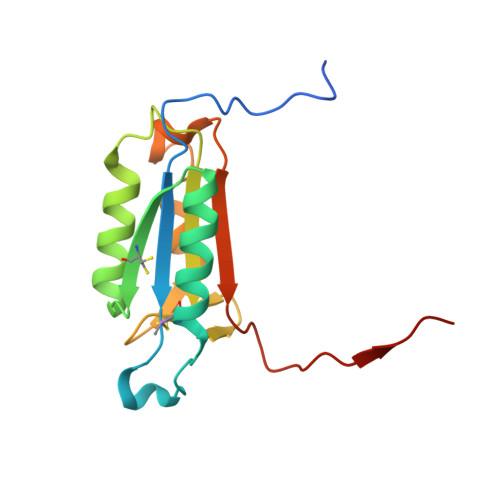 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P55212 GTEx: ENSG00000138794 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P55212 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Caspase-6 | 108 | Homo sapiens | Mutation(s): 0 Gene Names: CASP6, MCH2 EC: 3.4.22.59 | 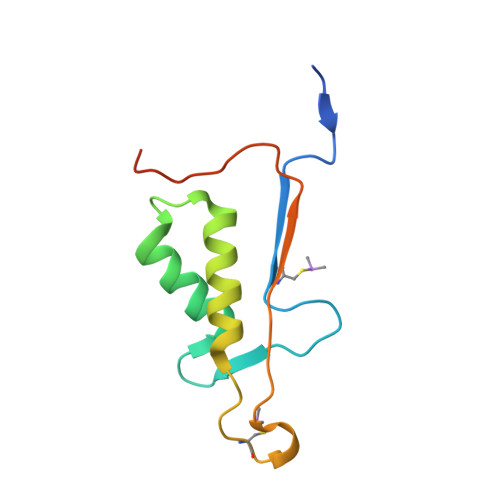 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P55212 GTEx: ENSG00000138794 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P55212 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| CAS Query on CAS | A, C | L-PEPTIDE LINKING | C5 H12 As N O2 S |  | CYS |
| ASJ Query on ASJ | E, F | PEPTIDE-LIKE | C4 H9 N O3 |  | -- |
| Entity ID: 3 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_000929 Query on PRD_000929 | E, F | N-acetyl-L-valyl-L-alpha-glutamyl-N-[(2S)-1-carboxy-3-hydroxypropan-2-yl]-L-isoleucinamide | Peptide-like / Inhibitor |  | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 55.455 | α = 90 |
| b = 89.5 | β = 111.73 |
| c = 61.245 | γ = 90 |
| Software Name | Purpose |
|---|---|
| MOSFLM | data reduction |
| SCALA | data scaling |
| MOLREP | phasing |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| CrystalClear | data collection |