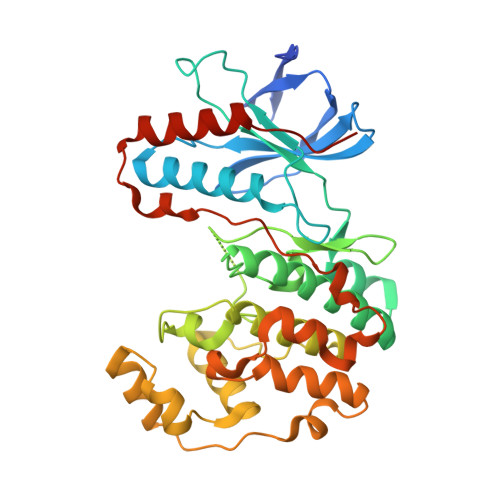
The third conformation of p38a MAP kinase observed in phosphorylated p38a and in solution
Akella, R., Min, X., Wu, Q., Gardner, K.H., Goldsmith, E.J.(2010) Structure 18: 1571-1578
- PubMed: 21134636 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2010.09.015
- Primary Citation Related Structures:
3P4K - PubMed Abstract:
MAPKs engage substrates, MAP2Ks, and phosphatases via a docking groove in the C-terminal domain of the kinase. Prior crystallographic studies on the unphosphorylated MAPKs p38α and ERK2 defined the docking groove and revealed long-range conformational changes affecting the activation loop and active site of the kinase induced by peptide. Solution NMR data presented here for unphosphorylated p38α with a MEK3b-derived peptide (p38α/pepMEK3b) validate these findings. Crystallograhic data from doubly phosphorylated active p38α (p38α/T∗GY∗/pepMEK3b) reveal a structure similar to unphosphorylated p38α/MEK3b, and distinct from phosphorylated p38γ (p38γ/T∗GY∗) and ERK2 (ERK2/T∗EY∗). The structure supports the idea that MAP kinases adopt three distinct conformations: unphosphorylated, phosphorylated, and a docking peptide-induced form.
- Department of Biochemistry, The University of Texas Southwestern Medical Center at Dallas, 5323 Harry Hines Boulevard, Dallas, TX 75390-8816, USA.
Organizational Affiliation: