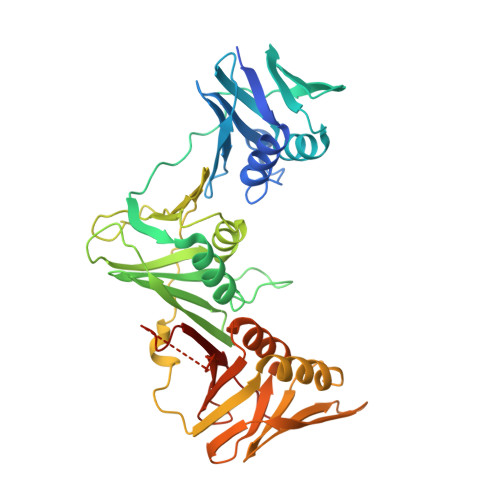
Crystal structure of DNA polymerase III beta sliding clamp from Mycobacterium tuberculosis.
Gui, W.J., Lin, S.Q., Chen, Y.Y., Zhang, X.E., Bi, L.J., Jiang, T.(2011) Biochem Biophys Res Commun 405: 272-277
- PubMed: 21219854 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2011.01.027
- Primary Citation Related Structures:
3P16 - PubMed Abstract:
The sliding clamp is a key component of DNA polymerase III (Pol III) required for genome replication. It is known to function with diverse DNA repair proteins and cell cycle-control proteins, making it a potential drug target. To extend our understanding of the structure/function relationship of the sliding clamp, we solved the crystal structure of the sliding clamp from Mycobacterium tuberculosis (M. tuberculosis), a human pathogen that causes most cases of tuberculosis (TB). The sliding clamp from M. tuberculosis forms a ring-shaped head-to-tail dimer with three domains per subunit. Each domain contains two α helices in the inner ring that lie against two β sheets in the outer ring. Previous studies have indicated that many Escherichia coli clamp-binding proteins have a conserved LF sequence, which is critical for binding to the hydrophobic region of the sliding clamp. Here, we analyzed the binding affinities of the M. tuberculosis sliding clamp and peptides derived from the α and δ subunits of Pol III, which indicated that the LF motif also plays an important role in the binding of the α and δ subunits to the sliding clamp of M. tuberculosis.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing 100101, China.
Organizational Affiliation: