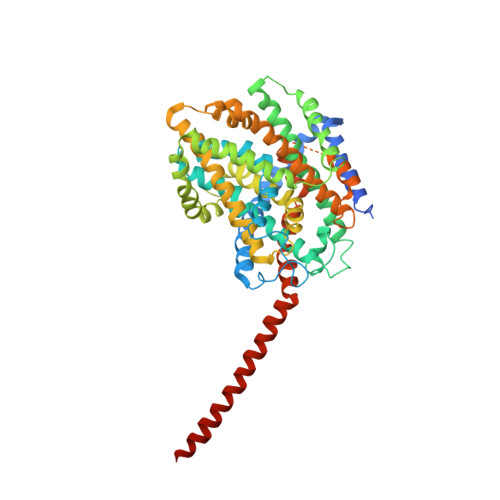
Substrate specificity and ion coupling in the Na(+)/betaine symporter BetP.
Perez, C., Koshy, C., Ressl, S., Nicklisch, S., Kramer, R., Ziegler, C.(2011) EMBO J 30: 1221-1229
- PubMed: 21364531 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.46
- Primary Citation Related Structures:
3P03 - PubMed Abstract:
BetP is an Na(+)-coupled betaine-specific transporter of the betaine-choline-carnitine (BCC) transporter family involved in the response to hyperosmotic stress. The crystal structure of BetP revealed an overall fold of two inverted structurally related repeats (LeuT-fold) that BetP shares with other sequence-unrelated Na(+)-coupled symporters. Numerous structures of LeuT-fold transporters in distinct conformational states have contributed substantially to our understanding of the alternating access mechanism of transport. Nevertheless, coupling of substrate and co-transported ion fluxes has not been structurally corroborated to the same extent. We converted BetP by a single-point mutation--glycine to aspartate--into an H(+)-coupled choline-specific transporter and solved the crystal structure of this mutant in complex with choline. The structure of BetP-G153D demonstrates a new inward-facing open conformation for BetP. Choline binding to a location close to the second, low-affinity sodium-binding site (Na2) of LeuT-fold transporters is facilitated by the introduced aspartate. Our data confirm the importance of a cation-binding site in BetP, playing a key role in a proposed molecular mechanism of Na(+) and H(+) coupling in BCC transporters.
- Department of Structural Biology, Max-Planck Institute of Biophysics, Frankfurt am Main, Germany.
Organizational Affiliation: