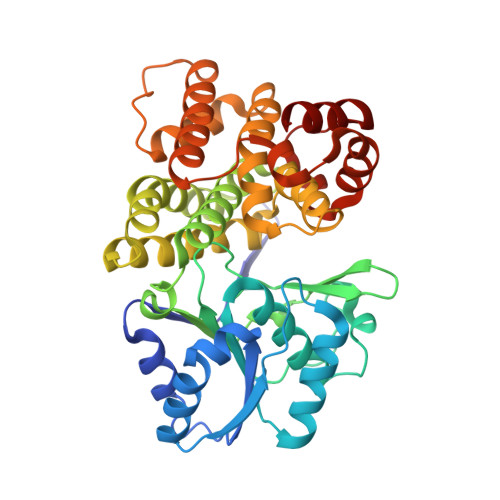
Structures of iron-dependent alcohol dehydrogenase 2 from Zymomonas mobilis ZM4 with and without NAD+ cofactor
Moon, J.H., Lee, H.J., Park, S.Y., Song, J.M., Park, M.Y., Park, H.M., Sun, J., Park, J.H., Kim, B.Y., Kim, J.S.(2011) J Mol Biology 407: 413-424
- PubMed: 21295587 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.01.045
- Primary Citation Related Structures:
3OWO, 3OX4 - PubMed Abstract:
The ethanologenic bacterium Zymomonas mobilis ZM4 is of special interest because it has a high ethanol yield. This is made possible by the two alcohol dehydrogenases (ADHs) present in Z. mobilis ZM4 (zmADHs), which shift the equilibrium of the reaction toward the synthesis of ethanol. They are metal-dependent enzymes: zinc for zmADH1 and iron for zmADH2. However, zmADH2 is inactivated by oxygen, thus implicating zmADH2 as the component of the cytosolic respiratory system in Z. mobilis. Here, we show crystal structures of zmADH2 in the form of an apo-enzyme and an NAD+–cofactor complex. The overall folding of the monomeric structure is very similar to those of other functionally related ADHs with structural variations around the probable substrate and NAD+ cofactor binding region. A dimeric structure is formed by the limited interactions between the two subunits with the bound NAD+ at the cleft formed along the domain interface. The catalytic iron ion binds near to the nicotinamide ring of NAD+, which is likely to restrict and locate the ethanol to the active site together with the oxidized Cys residue and several nonpolar bulky residues. The structures of the zmADH2 from the proficient ethanologenic bacterium Z. mobilis, with and without NAD+ cofactor, and modeling ethanol in the active site imply that there is a typical metal-dependent catalytic mechanism.
- Department of Chemistry and Institute of Basic Sciences, Chonnam National University, 300, Yongbong-dong, Buk-gu, Gwangju 500-757, Korea.
Organizational Affiliation: