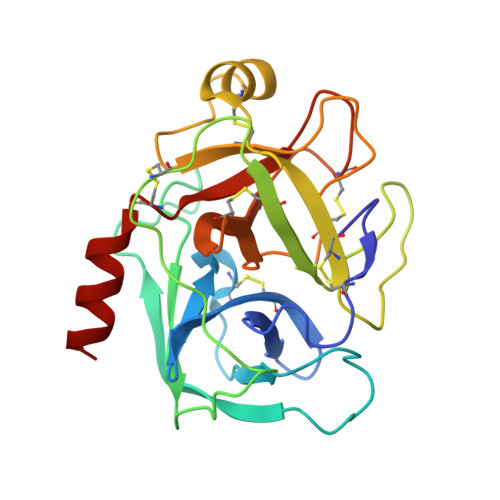
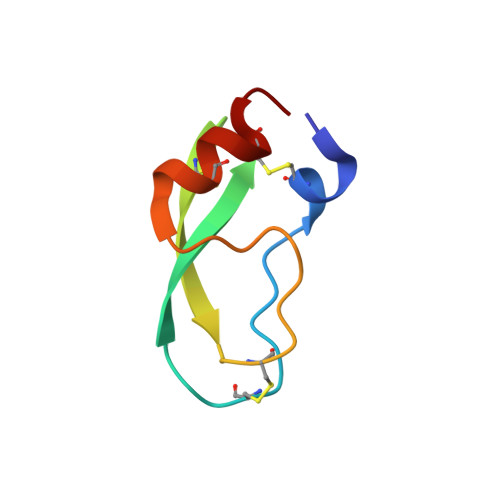
X-ray and neutron protein crystallographic analysis of the trypsin-BPTI complex.
Kawamura, K., Yamada, T., Kurihara, K., Tamada, T., Kuroki, R., Tanaka, I., Takahashi, H., Niimura, N.(2011) Acta Crystallogr D Biol Crystallogr 67: 140-148
- PubMed: 21245536 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444910053382
- Primary Citation Related Structures:
3OTJ - PubMed Abstract:
In this work, the crystal structure of the β-trypsin-bovine pancreatic trypsin inhibitor (BPTI) complex was refined and the D and H atoms in the complex were identified using data from both 1.6 Å resolution X-ray diffraction and 2.15 Å resolution neutron diffraction. After crystallization in an H(2)O solution, the sample crystal was soaked in a D(2)O solution for about two weeks. The protonation states of the catalytic triad (Asp102, His57 and Ser195) were observed. These results confirmed that the nucleophilic reactivity of the hydroxyl group of Ser195 was increased by forming a hydrogen bond with His57. According to structural analysis, the trypsin-BPTI interfaces located at the scissile peptide and the active sites were inaccessible to solvent water, and the amide H atoms of P2' Arg17/I, Gly216/E and Gly193/E at the binding interface were protected from H/D exchange. In contrast, both the amide H atom of P1' Ala16/I of the scissile peptide bond P1-P1' and the H atom between His57 N(ℇ2) and Ser195 O(γ) were replaced by D atoms. The hydrogen-bond networks at the S1 pocket were also confirmed and discussed from the viewpoint of substrate recognition. Moreover, the first neutron crystallographic structure of the Michaelis complex state of trypsin-BPTI is presented.
- Institute of Applied Beam Science, Graduate School of Science and Engineering, Ibaraki University, 4-12-1 Naka-Narusawa, Hitachi, Ibaraki 316-8511, Japan.
Organizational Affiliation: