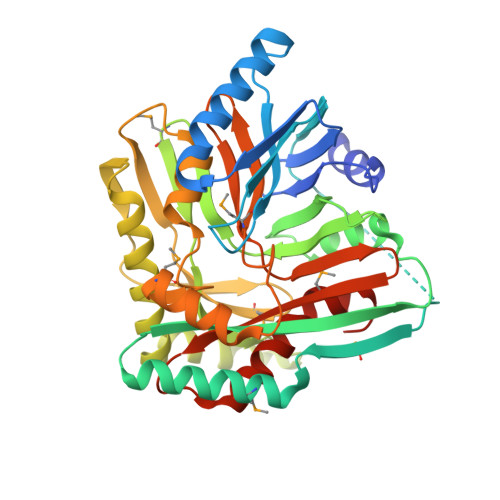
Structure of isochorismate synthase DhbC from Bacillus anthracis.
Domagalski, M.J., Tkaczuk, K.L., Chruszcz, M., Skarina, T., Onopriyenko, O., Cymborowski, M., Grabowski, M., Savchenko, A., Minor, W.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 956-961
- PubMed: 23989140 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113021246
- Primary Citation Related Structures:
3OS6 - PubMed Abstract:
The isochorismate synthase DhbC from Bacillus anthracis is essential for the biosynthesis of the siderophore bacillibactin by this pathogenic bacterium. The structure of the selenomethionine-substituted protein was determined to 2.4 Å resolution using single-wavelength anomalous diffraction. B. anthracis DhbC bears the strongest resemblance to the Escherichia coli isochorismate synthase EntC, which is involved in the biosynthesis of another siderophore, namely enterobactin. Both proteins adopt the characteristic fold of other chorismate-utilizing enzymes, which are involved in the biosynthesis of various products, including siderophores, menaquinone and tryptophan. The conservation of the active-site residues, as well as their spatial arrangement, suggests that these enzymes share a common Mg(2+)-dependent catalytic mechanism.
- Department of Molecular Physiology and Biological Physics, University of Virginia, 1340 Jefferson Park Avenue, Jordan Hall, Charlottesville, VA 22908, USA.
Organizational Affiliation: