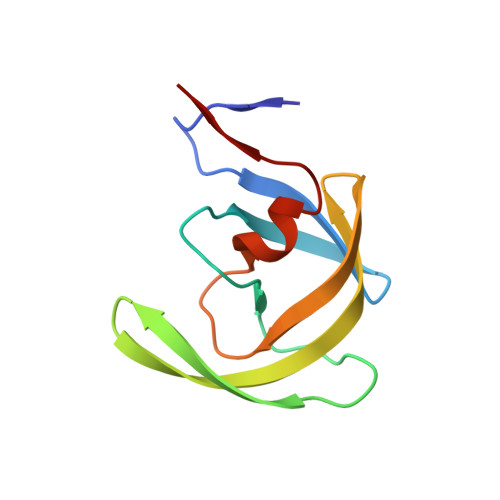
Probing Multidrug-Resistance and Protein-Ligand Interactions with Oxatricyclic Designed Ligands in HIV-1 Protease Inhibitors.
Ghosh, A.K., Xu, C.X., Rao, K.V., Baldridge, A., Agniswamy, J., Wang, Y.F., Weber, I.T., Aoki, M., Miguel, S.G., Amano, M., Mitsuya, H.(2010) ChemMedChem 5: 1850-1854
- PubMed: 20827746 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cmdc.201000318
- Primary Citation Related Structures:
3OK9 - Department of Chemistry and Medicinal Chemistry, Purdue University, 560 Oval Drive, West Lafayette, IN 47907, USA. akghosh@purdue.edu
Organizational Affiliation: