Synthesis and SAR of indole-and 7-azaindole-1,3-dicarboxamide hydroxyethylamine inhibitors of BACE-1.
Marcin, L.R., Higgins, M.A., Zusi, F.C., Zhang, Y., Dee, M.F., Parker, M.F., Muckelbauer, J.K., Camac, D.M., Morin, P.E., Ramamurthy, V., Tebben, A.J., Lentz, K.A., Grace, J.E., Marcinkeviciene, J.A., Kopcho, L.M., Burton, C.R., Barten, D.M., Toyn, J.H., Meredith, J.E., Albright, C.F., Bronson, J.J., Macor, J.E., Thompson, L.A.(2011) Bioorg Med Chem Lett 21: 537-541
- PubMed: 21078556 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2010.10.079
- Primary Citation Related Structures:
3OHF, 3OHH - PubMed Abstract:
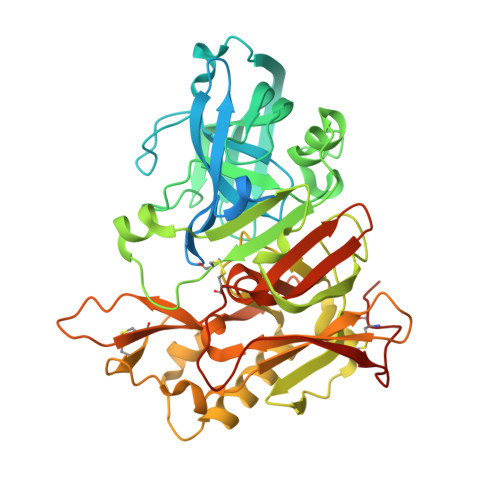
Heterocyclic replacement of the isophthalamide phenyl ring in hydroxyethylamine (HEA) BACE-1 inhibitors was explored. A variety of indole-1,3-dicarboxamide HEAs exhibited potent BACE-1 enzyme inhibition, but displayed poor cellular activity. Improvements in cellular activity and aspartic protease selectivity were observed for 7-azaindole-1,3-dicarboxamide HEAs. A methylprolinol-bearing derivative (10n) demonstrated robust reductions in rat plasma Aβ levels, but did not lower rat brain Aβ due to poor central exposure. The same analog exhibited a high efflux ratio in a bidirectional Caco-2 assay and was likely a substrate of the efflux transporter P-glycoprotein. X-ray crystal structures are reported for two indole HEAs in complex with BACE-1.
- Bristol-Myers Squibb, Research and Development, 5 Research Parkway, Wallingford, CT 06492-7660, USA. lawrence.marcin@bms.com
Organizational Affiliation: