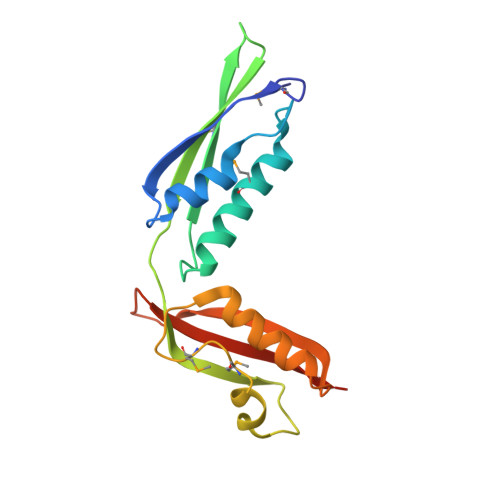
Structure and Flexibility of the Complete Periplasmic Domain of BamA: The Protein Insertion Machine of the Outer Membrane
Gatzeva-Topalova, P.Z., Warner, L.R., Pardi, A., Sousa, M.C.(2010) Structure 18: 1492-1501
- PubMed: 21070948 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2010.08.012
- Primary Citation Related Structures:
3OG5 - PubMed Abstract:
Folding and insertion of β-barrel outer membrane proteins (OMPs) is essential for Gram-negative bacteria. This process is mediated by the multiprotein complex BAM, composed of the essential β-barrel OMP BamA and four lipoproteins (BamBCDE). The periplasmic domain of BamA is key for its function and contains five "polypeptide transport-associated" (POTRA) repeats. Here, we report the crystal structure of the POTRA4-5 tandem, containing the essential for BAM complex formation and cell viability POTRA5. The domain orientation observed in the crystal is validated by solution NMR and SAXS. Using previously determined structures of BamA POTRA1-4, we present a spliced model of the entire BamA periplasmic domain validated by SAXS. Solution scattering shows that conformational flexibility between POTRA2 and 3 gives rise to compact and extended conformations. The length of BamA in its extended conformation suggests that the protein may bridge the inner and outer membranes across the periplasmic space.
- Department of Chemistry and Biochemistry, University of Colorado at Boulder, Boulder, CO 80309, USA.
Organizational Affiliation: