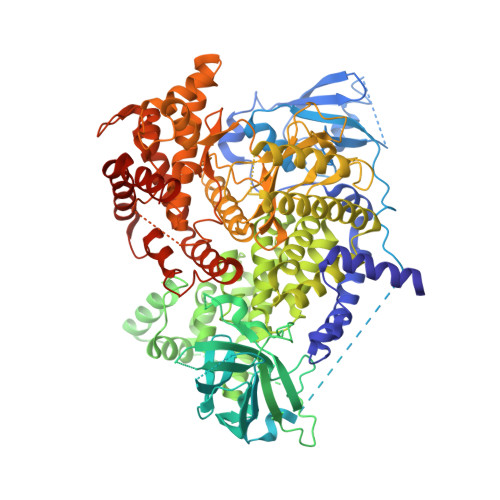
4-methylpteridinones as orally active and selective PI3K/mTOR dual inhibitors.
Liu, K.K., Bagrodia, S., Bailey, S., Cheng, H., Chen, H., Gao, L., Greasley, S., Hoffman, J.E., Hu, Q., Johnson, T.O., Knighton, D., Liu, Z., Marx, M.A., Nambu, M.D., Ninkovic, S., Pascual, B., Rafidi, K., Rodgers, C.M., Smith, G.L., Sun, S., Wang, H., Yang, A., Yuan, J., Zou, A.(2010) Bioorg Med Chem Lett 20: 6096-6099
- PubMed: 20817449 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2010.08.045
- Primary Citation Related Structures:
3OAW - PubMed Abstract:
Pteridinones were designed based on a non-selective kinase template. Because of the uniqueness of the PI3K and mTOR binding pockets, a methyl group was introduced to C-4 position of the peteridinone core to give compounds with excellent selectivity for PI3K and mTOR. This series of compounds were further optimized to improve their potency against PI3Kα and mTOR. Finally, orally active compounds with improved solubility and robust in vivo efficacy in tumor growth inhibition were identified as well.
- Pfizer Global Research and Development, Chemistry Department, La Jolla, CA 92120, USA. kevin.k.liu@pfizer.com
Organizational Affiliation: