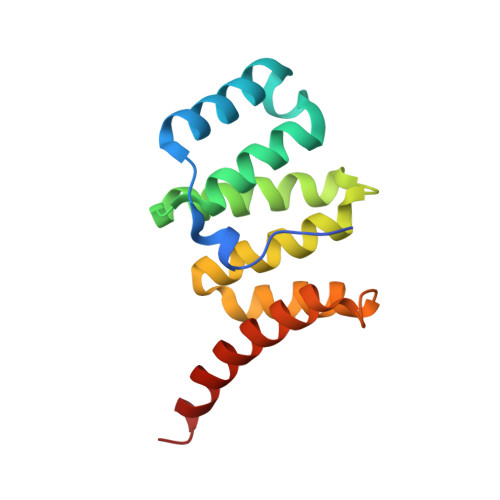
The 1.75 Angstrom resolution structure of fission protein Fis1 from Saccharomyces cerevisiae reveals elusive interactions of the autoinhibitory domain
Tooley, J.E., Khangulov, V., Lees, J.P., Schlessman, J.L., Bewley, M.C., Heroux, A., Bosch, J., Hill, R.B.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1310-1315
- PubMed: 22102223 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111029368
- Primary Citation Related Structures:
3O48 - PubMed Abstract:
Fis1 mediates mitochondrial and peroxisomal fission. It is tail-anchored to these organelles by a transmembrane domain, exposing a soluble cytoplasmic domain. Previous studies suggested that Fis1 is autoinhibited by its N-terminal region. Here, a 1.75 Å resolution crystal structure of the Fis1 cytoplasmic domain from Saccharomyces cerevisiae is reported which adopts a tetratricopeptide-repeat fold. It is observed that this fold creates a concave surface important for fission, but is sterically occluded by its N-terminal region. Thus, this structure provides a physical basis for autoinhibition and allows a detailed examination of the interactions that stabilize the inhibited state of this molecule.
- Department of Biology, Johns Hopkins University, Baltimore, MD 21218, USA.
Organizational Affiliation: