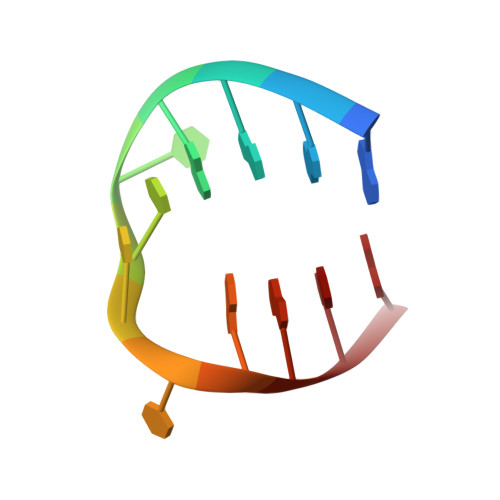
Fluorine in medicinal chemistry: beta-fluorination of peripheral pyrrolidines attached to acridine ligands affects their interactions with G-quadruplex DNA.
Campbell, N.H., Smith, D.L., Reszka, A.P., Neidle, S., O'Hagan, D.(2011) Org Biomol Chem 9: 1328-1331
- PubMed: 21221451 Search on PubMed
- DOI: https://doi.org/10.1039/c0ob00886a
- Primary Citation Related Structures:
3NYP, 3NZ7 - PubMed Abstract:
Comparative X-ray structure studies reveal that C-F bond incorporation into the peripheral pyrrolidine moieties of the G-quadruplex DNA binding ligand BSU6039 leads to a distinct pyrrolidine ring conformation, relative to the non-fluorinated analogue, and with a different binding mode involving reversal of the pyrrolidinium N(+)-H orientation.
- The School of Pharmacy, University of London, 29-39 Brunswick Square, London, UK WC1N 1AX.
Organizational Affiliation: