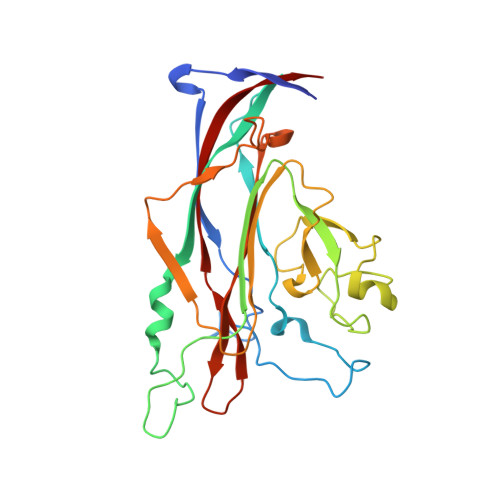
Structure-function analysis of the human JC polyomavirus establishes the LSTc pentasaccharide as a functional receptor motif.
Neu, U., Maginnis, M.S., Palma, A.S., Stroh, L.J., Nelson, C.D., Feizi, T., Atwood, W.J., Stehle, T.(2010) Cell Host Microbe 8: 309-319
- PubMed: 20951965 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chom.2010.09.004
- Primary Citation Related Structures:
3NXD, 3NXG - PubMed Abstract:
The human JC polyomavirus (JCV) causes a fatal demyelinating disease, progressive multifocal leukoencephalopathy (PML), in immunocompromised individuals. Current treatment options for PML are inadequate. Sialylated oligosaccharides and the serotonin receptor are known to be necessary for JCV entry, but the molecular interactions underlying JCV attachment remain unknown. Using glycan array screening and viral infectivity assays, we identify a linear sialylated pentasaccharide with the sequence NeuNAc-α2,6-Gal-β1,4-GlcNAc-β1,3-Gal-β1,4-Glc (LSTc) present on host glycoproteins and glycolipids as a specific JCV recognition motif. The crystal structure of the JCV capsid protein VP1 was solved alone and in complex with LSTc. It reveals extensive interactions with the terminal sialic acid of the LSTc motif and specific recognition of an extended conformation of LSTc. Mutations in the JCV oligosaccharide-binding sites abolish cell attachment, viral spread, and infectivity, further validating the importance of this interaction. Our findings provide a powerful platform for the development of antiviral compounds.
- Interfaculty Institute for Biochemistry, University of Tübingen, D-72076 Tübingen, Germany.
Organizational Affiliation: