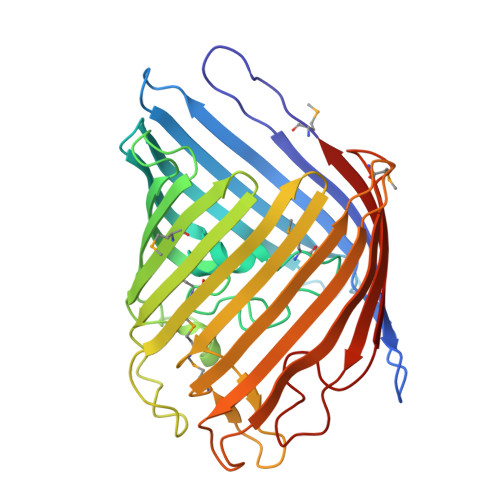
Asymmetric pore occupancy in crystal structure of OmpF porin from Salmonella typhi
Balasubramaniam, D., Arockiasamy, A., Kumar, P.D., Sharma, A., Krishnaswamy, S.(2012) J Struct Biol 178: 233-244
- PubMed: 22525817 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2012.04.005
- Primary Citation Related Structures:
3NSG - PubMed Abstract:
OmpF is a major general diffusion porin of Salmonella typhi, a Gram-negative bacterium, which is an obligatory human pathogen causing typhoid. The structure of S. typhi Ty21a OmpF (PDB Id: 3NSG) determined at 2.8 Å resolution by X-ray crystallography shows a 16-stranded β-barrel with three β-barrel monomers associated to form a trimer. The packing observed in S. typhi Ty21a rfOmpF crystals has not been observed earlier in other porin structures. The variations seen in the loop regions provide a starting point for using the S. typhi OmpF for structure-based multi-valent vaccine design. Along one side of the S. typhi Ty21a OmpF pore there exists a staircase arrangement of basic residues (20R, 60R, 62K, 65R, 77R, 130R and 16K), which also contribute, to the electrostatic potential in the pore. This structure suggests the presence of asymmetric electrostatics in the porin oligomer. Moreover, antibiotic translocation, permeability and reduced uptake in the case of mutants can be understood based on the structure paving the way for designing new antibiotics.
- Department of Genetic Engineering, School of Biotechnology, Madurai Kamaraj University, Madurai 625 021, India.
Organizational Affiliation: