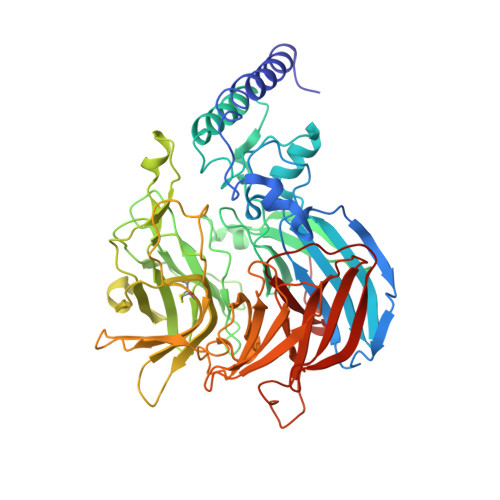
Structural insights into maize viviparous14, a key enzyme in the biosynthesis of the phytohormone abscisic acid.
Messing, S.A., Gabelli, S.B., Echeverria, I., Vogel, J.T., Guan, J.C., Tan, B.C., Klee, H.J., McCarty, D.R., Amzel, L.M.(2010) Plant Cell 22: 2970-2980
- PubMed: 20884803 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1105/tpc.110.074815
- Primary Citation Related Structures:
3NPE - PubMed Abstract:
The key regulatory step in the biosynthesis of abscisic acid (ABA), a hormone central to the regulation of several important processes in plants, is the oxidative cleavage of the 11,12 double bond of a 9-cis-epoxycarotenoid. The enzyme viviparous14 (VP14) performs this cleavage in maize (Zea mays), making it a target for the rational design of novel chemical agents and genetic modifications that improve plant behavior through the modulation of ABA levels. The structure of VP14, determined to 3.2-Å resolution, provides both insight into the determinants of regio- and stereospecificity of this enzyme and suggests a possible mechanism for oxidative cleavage. Furthermore, mutagenesis of the distantly related CCD1 of maize shows how the VP14 structure represents a template for all plant carotenoid cleavage dioxygenases (CCDs). In addition, the structure suggests how VP14 associates with the membrane as a way of gaining access to its membrane soluble substrate.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University, School of Medicine, Baltimore, Maryland 21205, USA.
Organizational Affiliation: