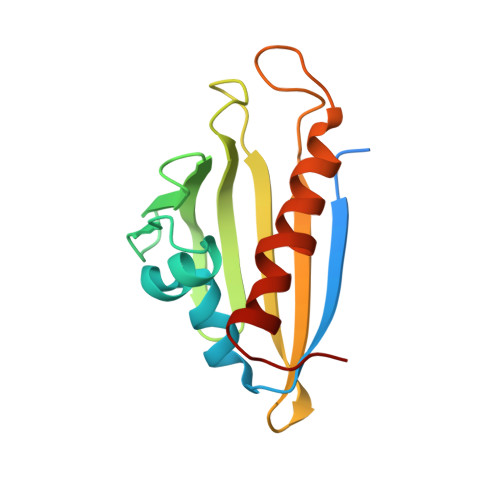
Crystal Structure of PFC0360w, an HSP90 activator from plasmodium falciparum
Wernimont, A.K., Hutchinson, A., Sullivan, H., MacKenzie, F., Kozieradzki, I., Cossar, D., Bochkarev, A., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Weigelt, J., Hui, R., Pizzaro, J.C., Hills, T.To be published.