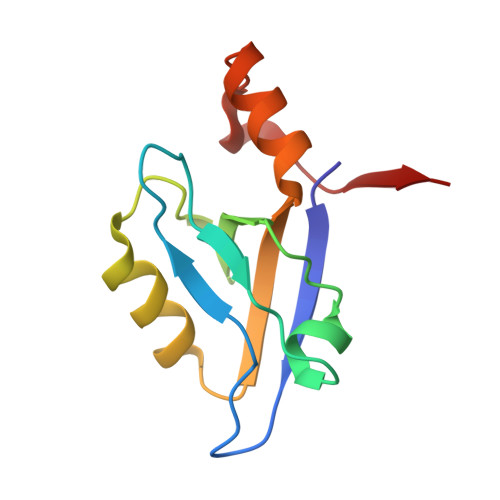
In vitro and in vivo analysis of the binding of the C terminus of the HDL receptor scavenger receptor class B, type I (SR-BI), to the PDZ1 domain of its adaptor protein PDZK1.
Kocher, O., Birrane, G., Tsukamoto, K., Fenske, S., Yesilaltay, A., Pal, R., Daniels, K., Ladias, J.A., Krieger, M.(2010) J Biological Chem 285: 34999-35010
- PubMed: 20739281 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.164418
- Primary Citation Related Structures:
3NGH - PubMed Abstract:
The PDZ1 domain of the four PDZ domain-containing protein PDZK1 has been reported to bind the C terminus of the HDL receptor scavenger receptor class B, type I (SR-BI), and to control hepatic SR-BI expression and function. We generated wild-type (WT) and mutant murine PDZ1 domains, the mutants bearing single amino acid substitutions in their carboxylate binding loop (Lys(14)-Xaa(4)-Asn(19)-Tyr-Gly-Phe-Phe-Leu(24)), and measured their binding affinity for a 7-residue peptide corresponding to the C terminus of SR-BI ((503)VLQEAKL(509)). The Y20A and G21Y substitutions abrogated all binding activity. Surprisingly, binding affinities (K(d)) of the K14A and F22A mutants were 3.2 and 4.0 μM, respectively, similar to 2.6 μM measured for the WT PDZ1. To understand these findings, we determined the high resolution structure of WT PDZ1 bound to a 5-residue sequence from the C-terminal SR-BI ((505)QEAKL(509)) using x-ray crystallography. In addition, we incorporated the K14A and Y20A substitutions into full-length PDZK1 liver-specific transgenes and expressed them in WT and PDZK1 knock-out mice. In WT mice, the transgenes did not alter endogenous hepatic SR-BI protein expression (intracellular distribution or amount) or lipoprotein metabolism (total plasma cholesterol, lipoprotein size distribution). In PDZK1 knock-out mice, as expected, the K14A mutant behaved like wild-type PDZK1 and completely corrected their hepatic SR-BI and plasma lipoprotein abnormalities. Unexpectedly, the 10-20-fold overexpressed Y20A mutant also substantially, but not completely, corrected these abnormalities. The results suggest that there may be an additional site(s) within PDZK1 that bind(s) SR-BI and mediate(s) productive SR-BI-PDZK1 interaction previously attributed exclusively to the canonical binding of the C-terminal SR-BI to PDZ1.
- Department of Pathology, Center for Vascular Biology Research, Division of Experimental Medicine, Beth Israel Deaconess Medical Center, Harvard Medical School, Boston, Massachusetts 02215, USA.okocher@bidmc.harvard.edu
Organizational Affiliation: