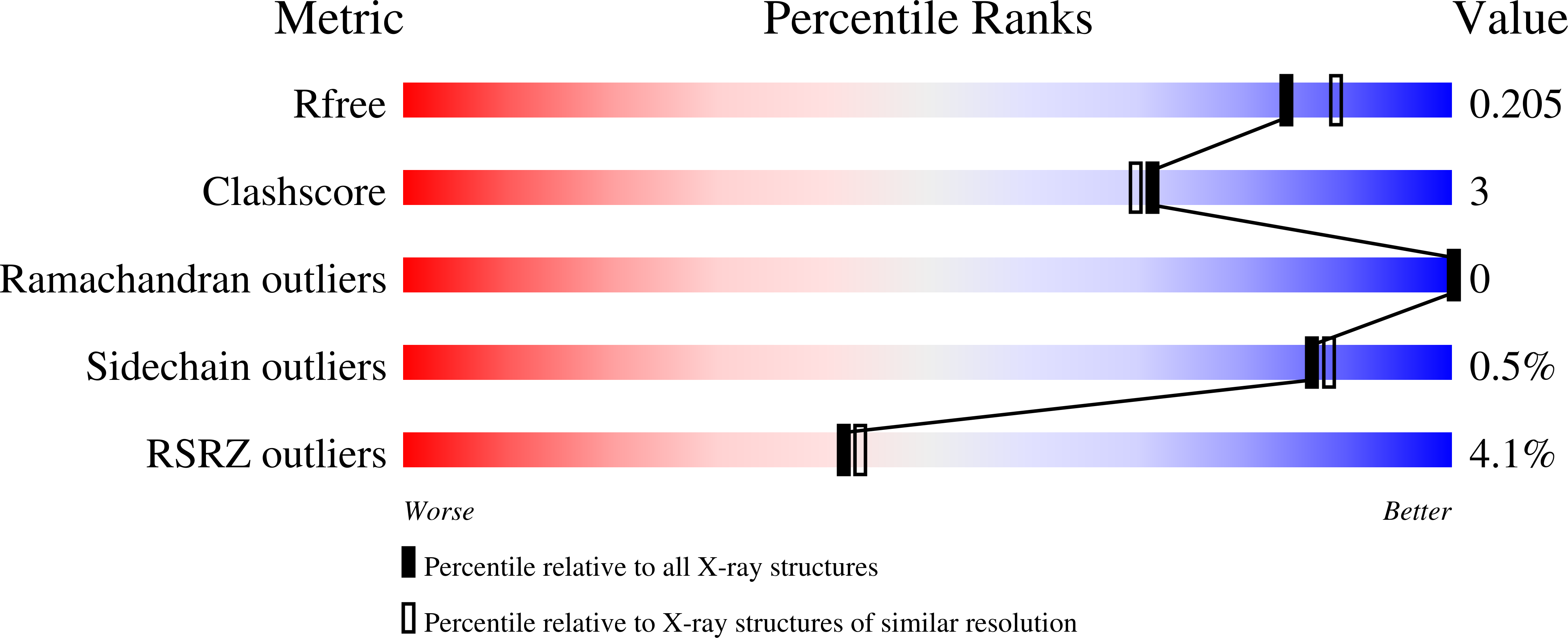
RNA Polymerase I Contains a TFIIF-Related DNA-Binding Subcomplex.
Geiger, S.R., Lorenzen, K., Schreieck, A., Hanecker, P., Kostrewa, D., Heck, A.J., Cramer, P.(2010) Mol Cell 39: 583-594
- PubMed: 20797630 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2010.07.028
- Primary Citation Related Structures:
3NFF, 3NFG, 3NFH, 3NFI - PubMed Abstract:
The eukaryotic RNA polymerases Pol I, II, and III use different promoters to transcribe different classes of genes. Promoter usage relies on initiation factors, including TFIIF and TFIIE, in the case of Pol II. Here, we show that the Pol I-specific subunits A49 and A34.5 form a subcomplex that binds DNA and is related to TFIIF and TFIIE. The N-terminal regions of A49 and A34.5 form a dimerization module that stimulates polymerase-intrinsic RNA cleavage and has a fold that resembles the TFIIF core. The C-terminal region of A49 forms a "tandem winged helix" (tWH) domain that binds DNA with a preference for the upstream promoter nontemplate strand and is predicted in TFIIE. Similar domains are predicted in Pol III-specific subunits. Thus, Pol I/III subunits that have no counterparts in Pol II are evolutionarily related to Pol II initiation factors and may have evolved to mediate promoter specificity and transcription processivity.
- Gene Center and Department of Biochemistry, Center for Integrated Protein Science CIPSM, Ludwig-Maximilians-Universität München, Feodor-Lynen-Strasse 25, 81377 Munich, Germany.
Organizational Affiliation: