The structure of the catalytic and carbohydrate binding domain of endoglucanase D bound to cellotriose
Bianchetti, C.M., Smith, R.W., Bingman, C.A., Phillips Jr., G.N.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Endoglucanase D | A, C [auth B], E [auth C], G [auth D] | 345 | Clostridium cellulovorans | Mutation(s): 0 Gene Names: engD EC: 3.2.1.4 | 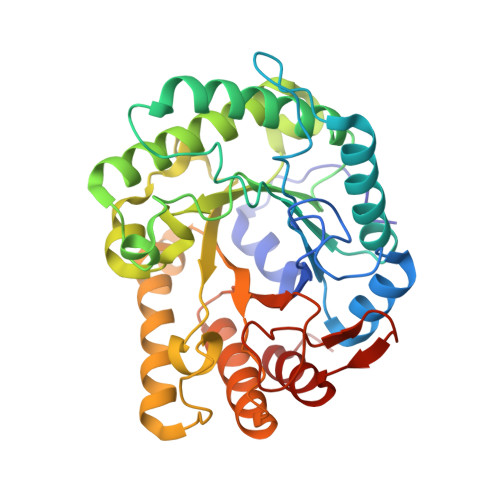 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P28623 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Endoglucanase D | B [auth E], D [auth F], F [auth G], H | 107 | Clostridium cellulovorans | Mutation(s): 0 Gene Names: engD EC: 3.2.1.4 | 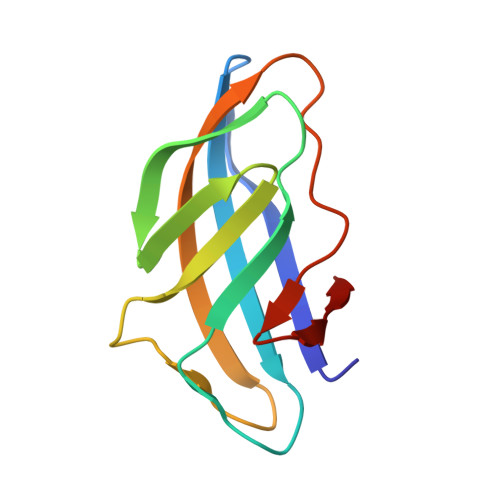 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P28623 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_900021 Query on PRD_900021 | I, J, K, L, M | beta-cellotriose | Oligosaccharide / Metabolism |  |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 85.297 | α = 90 |
| b = 119.05 | β = 90 |
| c = 198.487 | γ = 90 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| PHENIX | phasing |