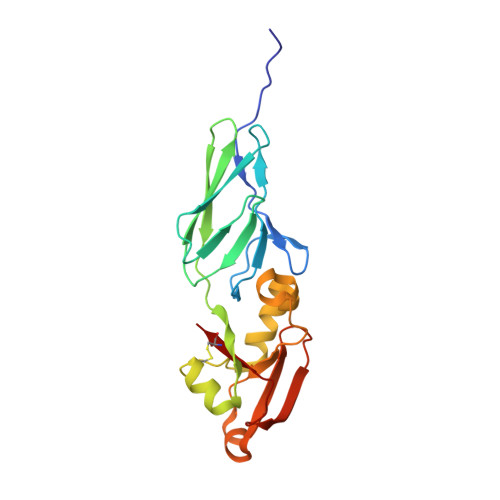
Crystal Structure of EHEC Intimin: Insights into the Complementarity between EPEC and EHEC
Yi, Y., Ma, Y., Gao, F., Mao, X., Peng, H., Feng, Y., Fan, Z., Wang, G., Guo, G., Yan, J., Zeng, H., Zou, Q.M., Gao, G.F.(2010) PLoS One 5: e15285-e15285
- PubMed: 21179574 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0015285
- Primary Citation Related Structures:
3NCW, 3NCX - PubMed Abstract:
Enterohaemorrhagic E. coli (EHEC) O157:H7 is a primary food-borne bacterial pathogen capable of causing life-threatening human infections which poses a serious challenge to public health worldwide. Intimin, the bacterial outer-membrane protein, plays a key role in the initiating process of EHEC infection. This activity is dependent upon translocation of the intimin receptor (Tir), the intimin binding partner of the bacteria-encoded host cell surface protein. Intimin has attracted considerable attention due to its potential function as an antibacterial drug target. Here, we report the crystal structure of the Tir-binding domain of intimin (Int188) from E. coli O157:H7 at 2.8 Å resolution, together with a mutant (IntN916Y) at 2.6 Å. We also built the structural model of EHEC intimin-Tir complex and analyzed the key binding residues. It suggested that the binding pattern of intimin and Tir between EHEC and Enteropathogenic E. coli (EPEC) adopt a similar mode and they can complement with each other. Detailed structural comparison indicates that there are four major points of structural variations between EHEC and EPEC intimins: one in Domain I (Ig-like domain), the other three located in Domain II (C-type lectin-like domain). These variations result in different binding affinities. These findings provide structural insight into the binding pattern of intimin to Tir and the molecular mechanism of EHEC O157: H7.
- Department of Clinical Microbiology and Clinical Immunology, College of Medical Laboratory, Third Military Medical University, Chongqing, China.
Organizational Affiliation: