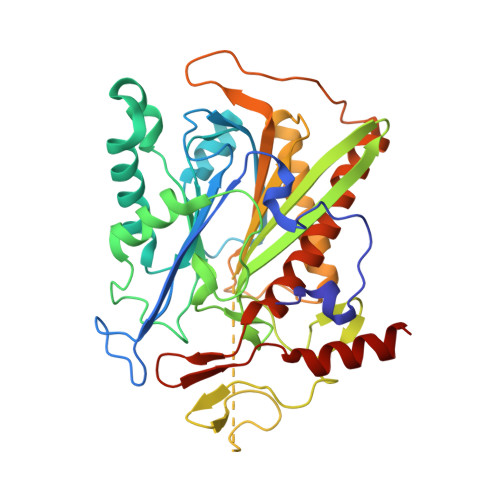
Autoproteolytic and catalytic mechanisms for the beta-aminopeptidase BapA--a member of the Ntn hydrolase family.
Merz, T., Heck, T., Geueke, B., Mittl, P.R., Briand, C., Seebach, D., Kohler, H.P., Grutter, M.G.(2012) Structure 20: 1850-1860
- PubMed: 22980995 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2012.07.017
- Primary Citation Related Structures:
3N2W, 3N33, 3N5I - PubMed Abstract:
The β-aminopeptidase BapA from Sphingosinicella xenopeptidilytica belongs to the N-terminal nucleophile (Ntn) hydrolases of the DmpA-like family and has the unprecedented property of cleaving N-terminal β-amino acid residues from peptides. We determined the crystal structures of the native (αβ)₄ heterooctamer and of the 153 kDa precursor homotetramer at a resolution of 1.45 and 1.8 Å, respectively. These structures together with mutational analyses strongly support mechanisms for autoproteolysis and catalysis that involve residues Ser250, Ser288, and Glu290. The autoproteolytic mechanism is different from the one so far described for Ntn hydrolases. The structures together with functional data also provide insight into the discriminating features of the active site cleft that determine substrate specificity.
- Biochemistry Institute, University of Zürich, Winterthurerstrasse 190, 8057 Zürich, Switzerland.
Organizational Affiliation: