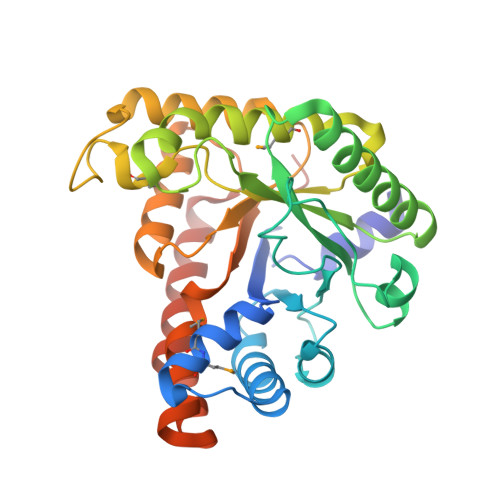
Crystal structures of LacD from Staphylococcus aureus and LacD.1 from Streptococcus pyogenes: Insights into substrate specificity and virulence gene regulation
Lee, S.J., Kim, H.S., Kim, D.J., Yoon, H.J., Kim, K.H., Yoon, J.Y., Suh, S.W.(2011) FEBS Lett 585: 307-312
- PubMed: 21192932 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2010.12.038
- Primary Citation Related Structures:
3MYO, 3MYP - PubMed Abstract:
Staphylococcus aureus LacD, a Class I tagatose-1,6-bisphosphate (TBP) aldolase, shows broadened substrate specificity by catalyzing the cleavage of 1,6-bisphosphate derivatives of D-tagatose, D-fructose, D-sorbose, and D-psicose. LacD.1 and LacD.2 are two closely-related Class I TBP aldolases in Streptococcus pyogenes. Here we have determined the crystal structures of S. aureus LacD and S. pyogenes LacD.1. Monomers of both enzymes are folded into a (β/α)(8) barrel and two monomers associate tightly to form a dimer in the crystals. The structures suggest that the residues E189 and S300 of rabbit muscle Class I fructose-1,6-bisphosphate (FBP) aldolase are important for substrate specificity. When we mutated the corresponding residues of S. aureus LacD, the mutants (L165E, L275S, and L165E/L275S) showed enhanced substrate specificity toward FBP.
- Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul, Republic of Korea.
Organizational Affiliation: