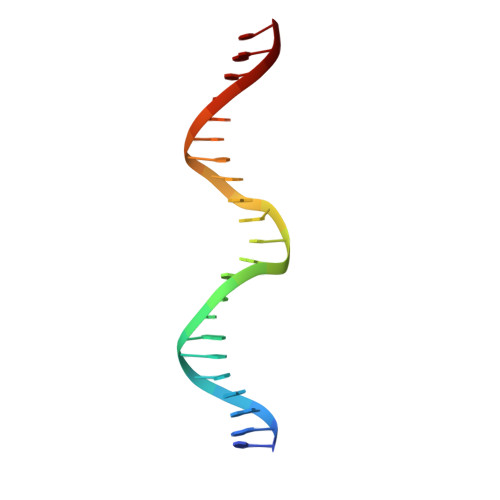
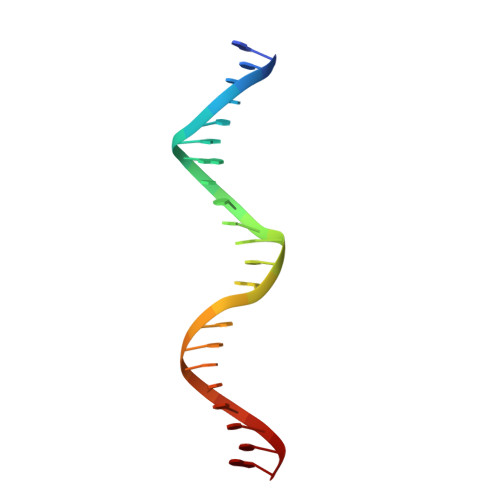
Folding, DNA Recognition, and Function of GIY-YIG Endonucleases: Crystal Structures of R.Eco29kI.
Mak, A.N., Lambert, A.R., Stoddard, B.L.(2010) Structure 18: 1321-1331
- PubMed: 20800503 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2010.07.006
- Primary Citation Related Structures:
3MX1, 3MX4, 3NIC - PubMed Abstract:
The GIY-YIG endonuclease family comprises hundreds of diverse proteins and a multitude of functions; none have been visualized bound to DNA. The structure of the GIY-YIG restriction endonuclease R.Eco29kI has been solved both alone and bound to its target site. The protein displays a domain-swapped homodimeric structure with several extended surface loops encircling the DNA. Only three side chains from each protein subunit contact DNA bases, two directly and one via a bridging solvent molecule. Both tyrosine residues within the GIY-YIG motif are positioned in the catalytic center near a putative nucleophilic water; the remainder of the active site resembles the HNH endonuclease family. The structure illustrates how the GIY-YIG scaffold has been adapted for the highly specific recognition of a DNA restriction site, in contrast to nonspecific DNA cleavage by GIY-YIG domains in homing endonucleases or structure-specific cleavage by DNA repair enzymes such as UvrC.
- Division of Basic Sciences, Fred Hutchinson Cancer Research Center, Seattle, WA 98109, USA.
Organizational Affiliation: