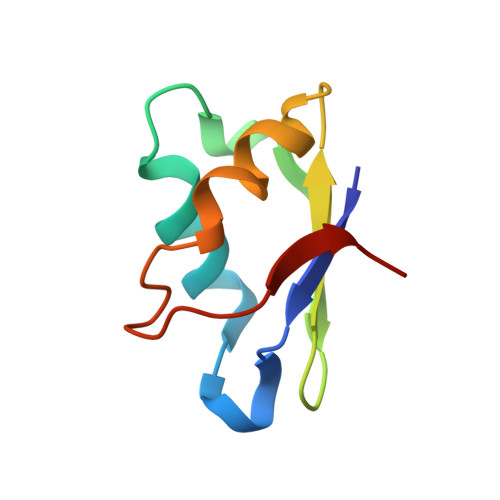
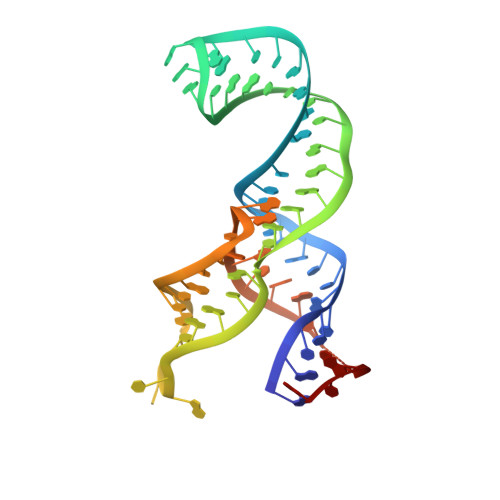
Structure of the RNA binding domain of a DEAD-box helicase bound to its ribosomal RNA target reveals a novel mode of recognition by an RNA recognition motif.
Hardin, J.W., Hu, Y.X., McKay, D.B.(2010) J Mol Biology 402: 412-427
- PubMed: 20673833 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.07.040
- Primary Citation Related Structures:
3MOJ - PubMed Abstract:
DEAD-box RNA helicases of the bacterial DbpA subfamily are localized to their biological substrate when a carboxy-terminal RNA recognition motif domain binds tightly and specifically to a segment of 23S ribosomal RNA (rRNA) that includes hairpin 92 of the peptidyl transferase center. A complex between a fragment of 23S rRNA and the RNA binding domain (RBD) of the Bacillus subtilis DbpA protein YxiN was crystallized and its structure was determined to 2.9 A resolution, revealing an RNA recognition mode that differs from those observed with other RNA recognition motifs. The RBD is bound between two RNA strands at a three-way junction. Multiple phosphates of the RNA backbone interact with an electropositive band generated by lysines of the RBD. Nucleotides of the single-stranded loop of hairpin 92 interact with the RBD, including the guanosine base of G2553, which forms three hydrogen bonds with the peptide backbone. A G2553U mutation reduces the RNA binding affinity by 2 orders of magnitude, confirming that G2553 is a sequence specificity determinant in RNA binding. Binding of the RBD to 23S rRNA in the late stages of ribosome subunit maturation would position the ATP-binding duplex destabilization fragment of the protein for interaction with rRNA in the peptidyl transferase cleft of the subunit, allowing it to "melt out" unstable secondary structures and allow proper folding.
- Department of Chemistry and Biochemistry, University of Colorado, Boulder, CO 80301, USA.
Organizational Affiliation: