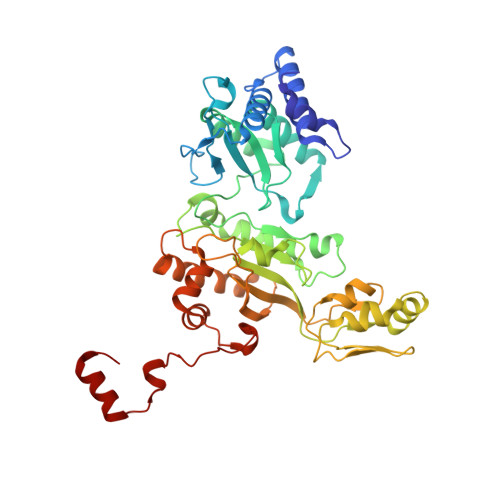
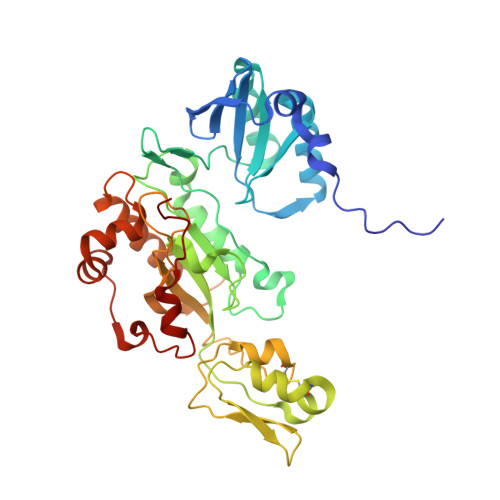
Structure of the dissimilatory sulfite reductase from the hyperthermophilic archaeon Archaeoglobus fulgidus.
Schiffer, A., Parey, K., Warkentin, E., Diederichs, K., Huber, H., Stetter, K.O., Kroneck, P.M., Ermler, U.(2008) J Mol Biology 379: 1063-1074
- PubMed: 18495156 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.04.027
- Primary Citation Related Structures:
3MMC - PubMed Abstract:
Conservation of energy based on the reduction of sulfate is of fundamental importance for the biogeochemical sulfur cycle. A key enzyme of this ancient anaerobic process is the dissimilatory sulfite reductase (dSir), which catalyzes the six-electron reduction of sulfite to hydrogen sulfide under participation of a unique magnetically coupled siroheme-[4Fe-4S] center. We determined the crystal structure of the enzyme from the sulfate-reducing archaeon Archaeoglobus fulgidus at 2-A resolution and compared it with that of the phylogenetically related assimilatory Sir (aSir). dSir is organized as a heterotetrameric (alphabeta)(2) complex composed of two catalytically independent alphabeta heterodimers. In contrast, aSir is a monomeric protein built of two fused modules that are structurally related to subunits alpha and beta except for a ferredoxin domain inserted only into the subunits of dSir. The [4Fe-4S] cluster of this ferredoxin domain is considered as the terminal redox site of the electron transfer pathway to the siroheme-[4Fe-4S] center in dSir. While aSir binds one siroheme-[4Fe-4S] center, dSir harbors two of them within each alphabeta heterodimer. Surprisingly, only one siroheme-[4Fe-4S] center in each alphabeta heterodimer is catalytically active, whereas access to the second one is blocked by a tryptophan residue. The spatial proximity of the functional and structural siroheme-[4Fe-4S] centers suggests that the catalytic activity at one active site was optimized during evolution at the expense of the enzymatic competence of the other. The sulfite binding mode and presumably the mechanism of sulfite reduction appear to be largely conserved between dSir and aSir. In addition, a scenario for the evolution of Sirs is proposed.
- Fachbereich Biologie, Mathematisch-Naturwissenschaftliche Sektion, Universität Konstanz, D-78457 Konstanz, Germany.
Organizational Affiliation: