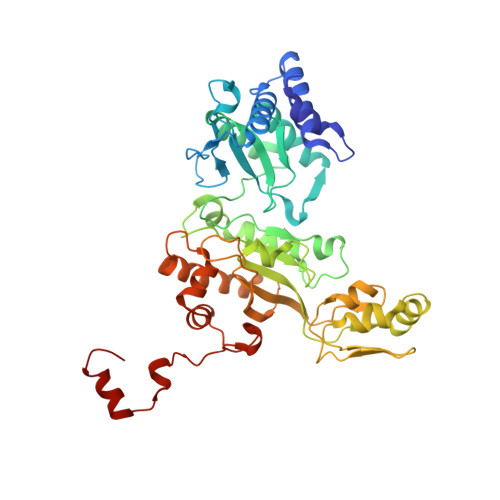
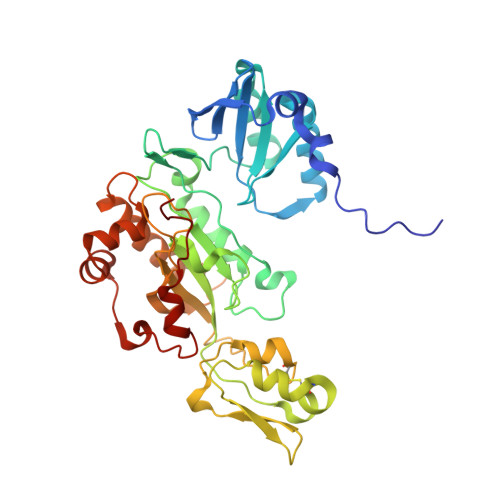
Reaction cycle of the dissimilatory sulfite reductase from Archaeoglobus fulgidus.
Parey, K., Warkentin, E., Kroneck, P.M., Ermler, U.(2010) Biochemistry 49: 8912-8921
- PubMed: 20822098 Search on PubMed
- DOI: https://doi.org/10.1021/bi100781f
- Primary Citation Related Structures:
3MM5, 3MM6, 3MM7, 3MM8, 3MM9, 3MMA, 3MMB - PubMed Abstract:
A vital process in the biogeochemical sulfur cycle is the dissimilatory sulfate reduction pathway in which sulfate (SO₄⁻²) is converted to hydrogen sulfide (H₂S). Dissimilatory sulfite reductase (dSir), its key enzyme, hosts a unique siroheme-[4Fe-4S] cofactor and catalyzes the six-electron reduction of sulfite (SO₃²⁻) to H₂S. To explore this reaction, we determined the X-ray structures of dSir from the archaeon Archaeoglobus fulgidus in complex with sulfite, sulfide (S²⁻) carbon monoxide (CO), cyanide (CN⁻), nitrite (NO₂⁻), nitrate (NO₃⁻), and phosphate (PO₄³⁻). Activity measurements indicated that dSir of A. fulgidus reduces, besides sulfite and nitrite, thiosulfate (S₂O₃²⁻) and trithionate (S₃O₆²⁻) and produces the latter two compounds besides sulfide. On this basis, a three-step mechanism was proposed, each step consisting of a two-electron transfer, a two-proton uptake, and a dehydration event. In comparison, the related active site structures of the assimilatory sulfite reductase (aSir)- and dSir-SO₃²⁻complexes reveal different conformations of Argα170 and Lysα211 both interacting with the sulfite oxygens (its sulfur atom coordinates the siroheme iron), a sulfite rotation of ~60° relative to each other, and different access of solvent molecules to the sulfite oxygens from the active site cleft. Therefore, solely in dSir a further sulfite molecule can be placed in van der Waals contact with the siroheme-ligated sulfite or sulfur-oxygen intermediates necessary for forming thiosulfate and trithionate. Although reported for dSir from several sulfate-reducing bacteria, the in vivo relevance of their formation is questionable.
- Max-Planck-Institut für Biophysik, Max-von-Laue-Strasse 3, Frankfurt, Germany.
Organizational Affiliation: