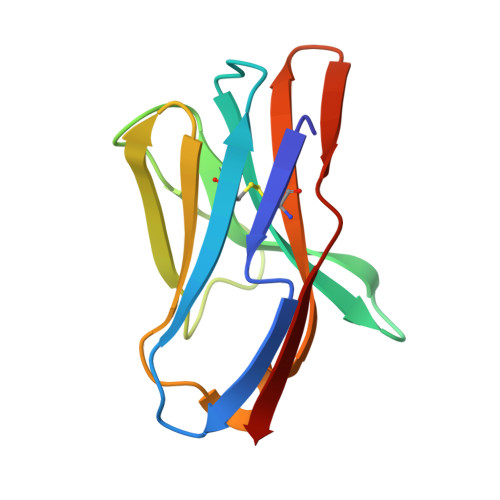
Crystal structure of staphylococcal enterotoxin G (SEG) in complex with a mouse T-cell receptor {beta} chain.
Fernandez, M.M., Cho, S., De Marzi, M.C., Kerzic, M.C., Robinson, H., Mariuzza, R.A., Malchiodi, E.L.(2011) J Biological Chem 286: 1189-1195
- PubMed: 21059660 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.142471
- Primary Citation Related Structures:
3MC0, 3OWE - PubMed Abstract:
Superantigens (SAgs) are bacterial or viral toxins that bind MHC class II (MHC-II) molecules and T-cell receptor (TCR) in a nonconventional manner, inducing T-cell activation that leads to inflammatory cytokine production, which may result in acute toxic shock. In addition, the emerging threat of purpura fulminans and community-associated meticillin-resistant Staphylococcus aureus emphasizes the importance of a better characterization of SAg binding to their natural ligands that may allow the development of reagents to neutralize their action. The three-dimensional structure of the complex between a mouse TCR β chain (mVβ8.2) and staphylococcal enterotoxin G (SEG) at 2.0 Å resolution revealed a binding site that does not conserve the "hot spots" present in mVβ8.2-SEC2, mVβ8.2-SEC3, mVβ8.2-SEB, and mVβ8.2-SPEA complexes. Analysis of the mVβ8.2-SEG interface allowed us to explain the higher affinity of this complex compared with the others, which may account for the early activation of T-cells bearing mVβ8.2 by SEG. This mode of interaction between SEG and mVβ8.2 could be an adaptive advantage to bestow on the pathogen a faster rate of colonization of the host.
- Cátedra de Inmunología and Instituto de Estudios de la Inmunidad Humoral Prof Ricardo A. Margni, Consejo Nacional de Investigaciones Científicas y Técnicas, Facultad de Farmacia y Bioquímica, Universidad de Buenos Aires, Junín 956 4to P, 1113 Buenos Aires, Argentina.
Organizational Affiliation: