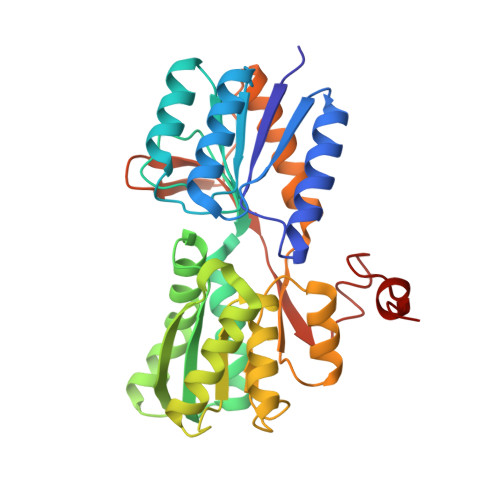
Conformational changes and ligand recognition of Escherichia coli D-xylose binding protein revealed
Sooriyaarachchi, S., Ubhayasekera, W., Park, C., Mowbray, S.L.(2010) J Mol Biology 402: 657-668
- PubMed: 20678502 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.07.038
- Primary Citation Related Structures:
3M9W, 3M9X, 3MA0 - PubMed Abstract:
ATP binding cassette transport systems account for most import of necessary nutrients in bacteria. The periplasmic binding component (or an equivalent membrane-anchored protein) is critical to recognizing cognate ligand and directing it to the appropriate membrane permease. Here we report the X-ray structures of D-xylose binding protein from Escherichia coli in ligand-free open form, ligand-bound open form, and ligand-bound closed form at 2.15 Å, 2.2 Å, and 2.2 Å resolutions, respectively. The ligand-bound open form is the first such structure to be reported at high resolution; the combination of the three different forms from the same protein furthermore gives unprecedented details concerning the conformational changes involved in binding protein function. As is typical of the structural family, the protein has two similar globular domains, which are connected by a three-stranded hinge region. The open liganded structure shows that xylose binds first to the C-terminal domain, with only very small conformational changes resulting. After a 34° closing motion, additional interactions are formed with the N-terminal domain; changes in this domain are larger and serve to make the structure more ordered near the ligand. An analysis of the interactions suggests why xylose is the preferred ligand. Furthermore, a comparison with the most closely related proteins in the structural family shows that the conformational changes are distinct in each type of binding protein, which may have implications for how the individual proteins act in concert with their respective membrane permeases.
- Department of Molecular Biology, Biomedical Center, Swedish University of Agricultural Sciences, Uppsala, Sweden.
Organizational Affiliation: