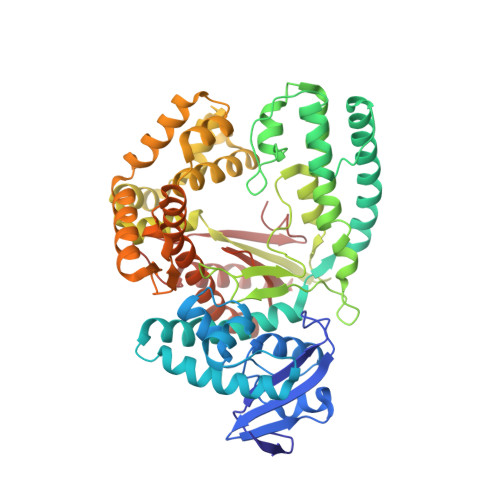
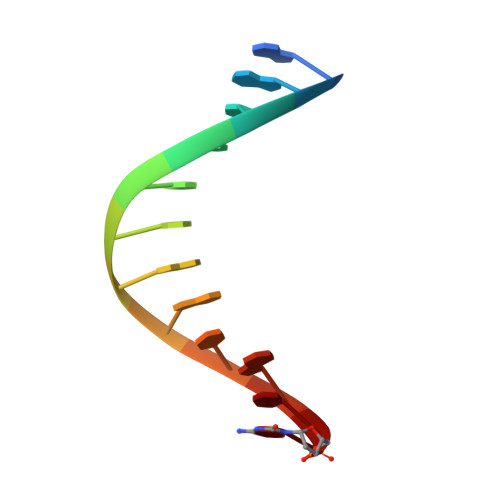
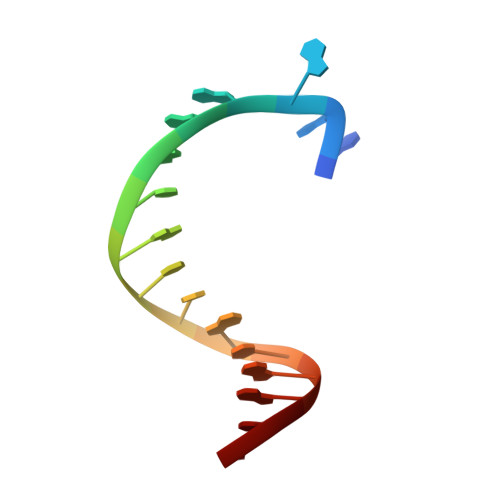
Structures of DNA polymerases caught processing size-augmented nucleotide probes.
Betz, K., Streckenbach, F., Schnur, A., Exner, T., Welte, W., Diederichs, K., Marx, A.(2010) Angew Chem Int Ed Engl 49: 5181-5184
- PubMed: 20572212 Search on PubMed
- DOI: https://doi.org/10.1002/anie.200905724
- Primary Citation Related Structures:
3M8R, 3M8S - Department of Chemistry, Konstanz Research School Chemical Biology, Universität Konstanz, Universitätsstrasse 10, 78457 Konstanz, Germany.
Organizational Affiliation: