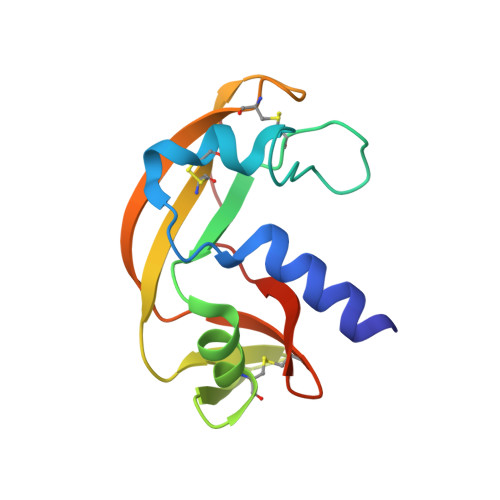
A new RNase sheds light on the RNase/angiogenin subfamily from zebrafish.
Pizzo, E., Merlino, A., Turano, M., Russo Krauss, I., Coscia, F., Zanfardino, A., Varcamonti, M., Furia, A., Giancola, C., Mazzarella, L., Sica, F., D'Alessio, G.(2010) Biochem J 433: 345-355
- PubMed: 21050179 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20100892
- Primary Citation Related Structures:
3LJD, 3LJE, 3LN8 - PubMed Abstract:
Recently, extracellular RNases of the RNase A superfamily, with the characteristic CKxxNTF sequence signature, have been identified in fish. This has led to the recognition that these RNases are present in the whole vertebrate subphylum. In fact, they comprise the only enzyme family unique to vertebrates. Four RNases from zebrafish (Danio rerio) have been previously reported and have a very low RNase activity; some of these are endowed, like human angiogenin, with powerful angiogenic and bactericidal activities. In the present paper, we report the three-dimensional structure, the thermodynamic behaviour and the biological properties of a novel zebrafish RNase, ZF-RNase-5. The investigation of its structural and functional properties, extended to all other subfamily members, provides an inclusive description of the whole zebrafish RNase subfamily.
- Dipartimento di Biologia Strutturale e Funzionale, Università di Napoli Federico II, Complesso Universitario Monte S. Angelo, Naples, Italy.
Organizational Affiliation: