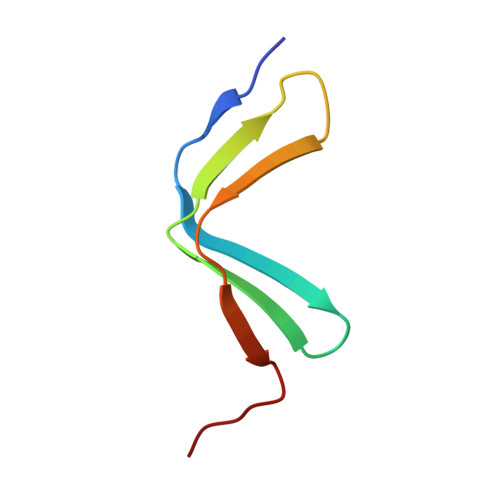
The crystal structure of the human nascent polypeptide-associated complex domain reveals a nucleic acid-binding region on the NACA subunit
Liu, Y., Hu, Y., Li, X., Niu, L., Teng, M.(2010) Biochemistry 49: 2890-2896
- PubMed: 20214399 Search on PubMed
- DOI: https://doi.org/10.1021/bi902050p
- Primary Citation Related Structures:
3LKX - PubMed Abstract:
In archaea and eukaryotes, the nascent polypeptide-associated complex (NAC) is one of the cytosolic chaperones that contact the nascent polypeptide chains as they emerge from the ribosome and assist in post-translational processes. The eukaryotic NAC is a heterodimer, and its two subunits form a stable complex through a dimerizing domain called the NAC domain. In addition to acting as a protein translation chaperone, the NAC subunits also function individually in transcriptional regulation. Here we report the crystal structure of the human NAC domain, which reveals the manner of human NAC dimerization. On the basis of the structure, we identified a region in the NAC domain of the human NAC alpha-subunit as a new nucleic acid-binding region, which is blocked from binding nucleic acids in the heterodimeric complex by a helix region in the beta-subunit.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230027, China.
Organizational Affiliation: