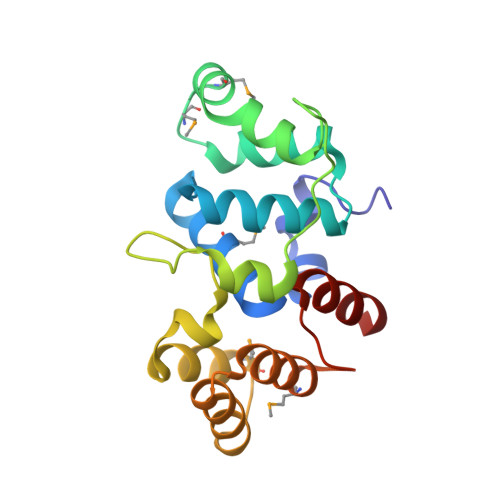
Shared Active Site Architecture between the Large Subunit of Eukaryotic Primase and DNA Photolyase
Sauguet, L., Klinge, S., Perera, R.L., Maman, J.D., Pellegrini, L.(2010) PLoS One 5: 10083-10083
- PubMed: 20404922 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0010083
- Primary Citation Related Structures:
3LGB - PubMed Abstract:
DNA synthesis during replication relies on RNA primers synthesised by the primase, a specialised DNA-dependent RNA polymerase that can initiate nucleic acid synthesis de novo. In archaeal and eukaryotic organisms, the primase is a heterodimeric enzyme resulting from the constitutive association of a small (PriS) and large (PriL) subunit. The ability of the primase to initiate synthesis of an RNA primer depends on a conserved Fe-S domain at the C-terminus of PriL (PriL-CTD). However, the critical role of the PriL-CTD in the catalytic mechanism of initiation is not understood. Here we report the crystal structure of the yeast PriL-CTD at 1.55 A resolution. The structure reveals that the PriL-CTD folds in two largely independent alpha-helical domains joined at their interface by a [4Fe-4S] cluster. The larger N-terminal domain represents the most conserved portion of the PriL-CTD, whereas the smaller C-terminal domain is largely absent in archaeal PriL. Unexpectedly, the N-terminal domain reveals a striking structural similarity with the active site region of the DNA photolyase/cryptochrome family of flavoproteins. The region of similarity includes PriL-CTD residues that are known to be essential for initiation of RNA primer synthesis by the primase. Our study reports the first crystallographic model of the conserved Fe-S domain of the archaeal/eukaryotic primase. The structural comparison with a cryptochrome protein bound to flavin adenine dinucleotide and single-stranded DNA provides important insight into the mechanism of RNA primer synthesis by the primase.
- Department of Biochemistry, University of Cambridge, Cambridge, United Kingdom.
Organizational Affiliation: