Unusual target site disruption by the rare-cutting HNH restriction endonuclease PacI.
Shen, B.W., Heiter, D.F., Chan, S.H., Wang, H., Xu, S.Y., Morgan, R.D., Wilson, G.G., Stoddard, B.L.(2010) Structure 18: 734-743
- PubMed: 20541511 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2010.03.009
- Primary Citation Related Structures:
3LDY, 3M7K - PubMed Abstract:
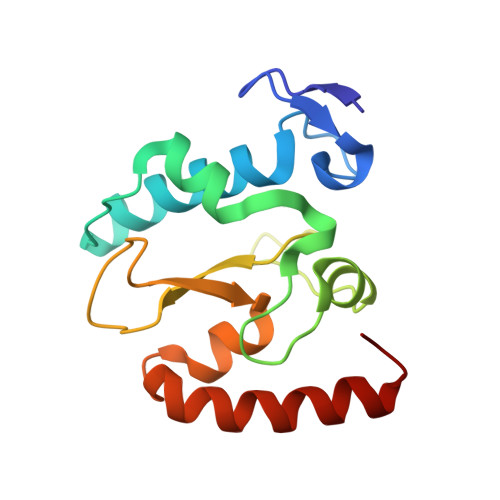
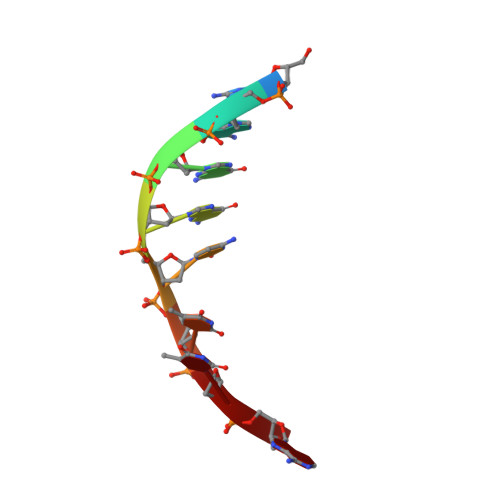
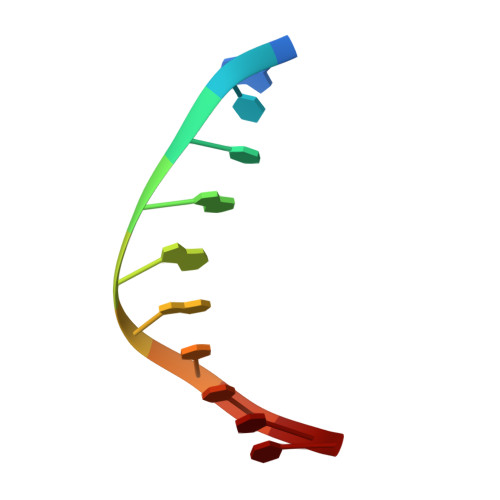
The crystal structure of the rare-cutting HNH restriction endonuclease PacI in complex with its eight-base-pair target recognition sequence 5'-TTAATTAA-3' has been determined to 1.9 A resolution. The enzyme forms an extended homodimer, with each subunit containing two zinc-bound motifs surrounding a betabetaalpha-metal catalytic site. The latter is unusual in that a tyrosine residue likely initiates strand cleavage. PacI dramatically distorts its target sequence from Watson-Crick duplex DNA base pairing, with every base separated from its original partner. Two bases on each strand are unpaired, four are engaged in noncanonical A:A and T:T base pairs, and the remaining two bases are matched with new Watson-Crick partners. This represents a highly unusual DNA binding mechanism for a restriction endonuclease, and implies that initial recognition of the target site might involve significantly different contacts from those visualized in the DNA-bound cocrystal structures.
- Division of Basic Sciences, Fred Hutchinson Cancer Research Center, 1100 Fairview Avenue N. A3-025, Seattle, WA 98109, USA.
Organizational Affiliation: