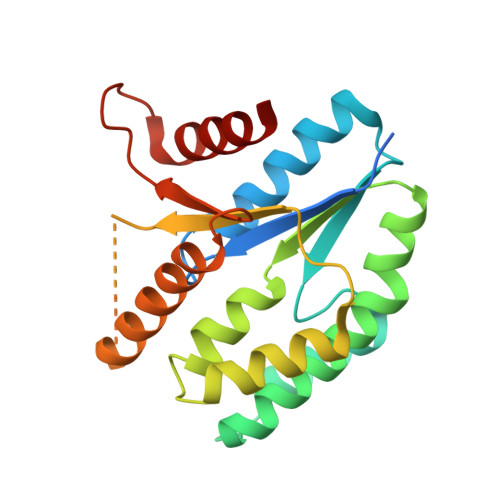
Structure of thymidylate kinase from Ehrlichia chaffeensis.
Leibly, D.J., Abendroth, J., Bryan, C.M., Sankaran, B., Kelley, A., Barrett, L.K., Stewart, L., Van Voorhis, W.C.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1090-1094
- PubMed: 21904055 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S174430911101493X
- Primary Citation Related Structures:
3LD9 - PubMed Abstract:
The enzyme thymidylate kinase phosphorylates the substrate thymidine 5'-phosphate (dTMP) to form thymidine 5'-diphosphate (dTDP), which is further phosphorylated to dTTP for incorporation into DNA. Ehrlichia chaffeensis is the etiologic agent of human monocytotropic erlichiosis (HME), a potentially life-threatening tick-borne infection. HME is endemic in the United States from the southern states up to the eastern seaboard. HME is transmitted to humans via the lone star tick Amblyomma americanum. Here, the 2.15 Å resolution crystal structure of thymidylate kinase from E. chaffeensis in the apo form is presented.
- Seattle Structural Genomics Center for Infectious Disease (SSGCID), USA.
Organizational Affiliation: