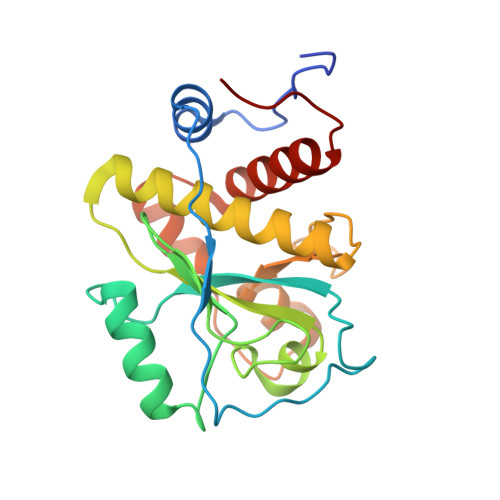
Structural basis of cell wall cleavage by a staphylococcal autolysin
Zoll, S., Patzold, B., Schlag, M., Gotz, F., Kalbacher, H., Stehle, T.(2010) PLoS Pathog 6: e1000807-e1000807
- PubMed: 20300605 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1000807
- Primary Citation Related Structures:
3LAT - PubMed Abstract:
The major autolysins (Atl) of Staphylococcus epidermidis and S. aureus play an important role in cell separation, and their mutants are also attenuated in virulence. Therefore, autolysins represent a promising target for the development of new types of antibiotics. Here, we report the high-resolution structure of the catalytically active amidase domain AmiE (amidase S. epidermidis) from the major autolysin of S. epidermidis. This is the first protein structure with an amidase-like fold from a bacterium with a gram-positive cell wall architecture. AmiE adopts a globular fold, with several alpha-helices surrounding a central beta-sheet. Sequence comparison reveals a cluster of conserved amino acids that define a putative binding site with a buried zinc ion. Mutations of key residues in the putative active site result in loss of activity, enabling us to propose a catalytic mechanism. We also identified and synthesized muramyltripeptide, the minimal peptidoglycan fragment that can be used as a substrate by the enzyme. Molecular docking and digestion assays with muramyltripeptide derivatives allow us to identify key determinants of ligand binding. This results in a plausible model of interaction of this ligand not only for AmiE, but also for other PGN-hydrolases that share the same fold. As AmiE active-site mutations also show a severe growth defect, our findings provide an excellent platform for the design of specific inhibitors that target staphylococcal cell separation and can thereby prevent growth of this pathogen.
- Interfaculty Institute for Biochemistry, University of Tübingen, Tübingen, Germany.
Organizational Affiliation: