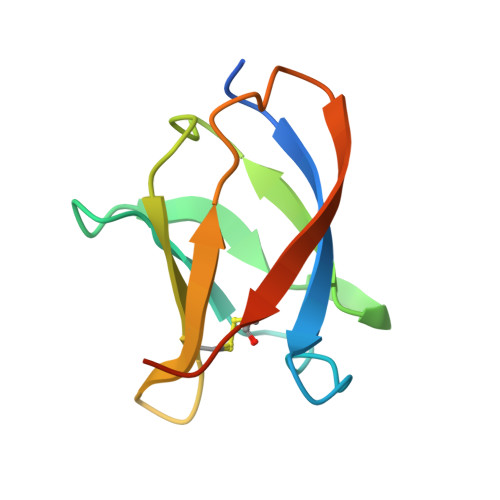
Structural Homology between the C-Terminal Domain of the PapC Usher and Its Plug.
Ford, B., Rego, A.T., Ragan, T.J., Pinkner, J., Dodson, K., Driscoll, P.C., Hultgren, S., Waksman, G.(2010) J Bacteriol 192: 1824-1831
- PubMed: 20118254 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.01677-09
- Primary Citation Related Structures:
2KT6, 3L48 - PubMed Abstract:
P pili are extracellular appendages responsible for the targeting of uropathogenic Escherichia coli to the kidney. They are assembled by the chaperone-usher (CU) pathway of pilus biogenesis involving two proteins, the periplasmic chaperone PapD and the outer membrane assembly platform, PapC. Many aspects of the structural biology of the Pap CU pathway have been elucidated, except for the C-terminal domain of the PapC usher, the structure of which is unknown. In this report, we identify a stable and folded fragment of the C-terminal region of the PapC usher and determine its structure using both X-ray crystallography and nuclear magnetic resonance (NMR) spectroscopy. These structures reveal a beta-sandwich fold very similar to that of the plug domain, a domain of PapC obstructing its translocation domain. This structural similarity suggests similar functions in usher-mediated pilus biogenesis, playing out at different stages of the process. This structure paves the way for further functional analysis targeting surfaces common to both the plug and the C-terminal domain of PapC.
- Department of Pathology and Immunology, Washington University School of Medicine, Saint Louis, Missouri 63110, USA.
Organizational Affiliation: