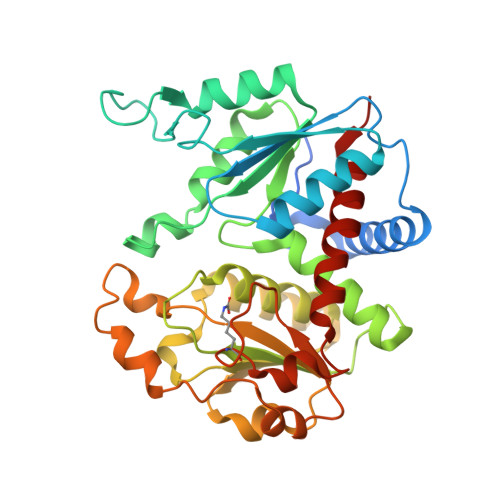
Crystal structure of N-acetylornithine transcarbamylase from Xanthomonas campestris: a novel enzyme in a new arginine biosynthetic pathway found in several eubacteria.
Shi, D., Morizono, H., Yu, X., Roth, L., Caldovic, L., Allewell, N.M., Malamy, M.H., Tuchman, M.(2005) J Biological Chem 280: 14366-14369
- PubMed: 15731101 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.C500005200
- Primary Citation Related Structures:
3KZC, 3KZK - PubMed Abstract:
We have identified in Xanthomonas campestris a novel N-acetylornithine transcarbamylase that replaces ornithine transcarbamylase in the canonic arginine biosynthetic pathway of several Eubacteria. The crystal structures of the protein in the presence and absence of the reaction product, N-acetylcitrulline, were determined. This new family of transcarbamylases lacks the DxxSMG motif that is characteristic of all ornithine transcarbamylases (OTCases) and contains a novel proline-rich loop that forms part of the active site. The specificity for N-acetylornithine is conferred by hydrogen bonding with residues in the proline-rich loop via water molecules and by hydrophobic interactions with residues from the adjacent 80's, 120's, and proline-rich loops. This novel protein structure provides a starting point for rational design of specific analogs that may be useful in combating human and plant pathogens that utilize acetylornithine transcarbamylase rather than ornithine transcarbamylase.
- Children's Research Institute, Children's National Medical Center, Washington, DC 20010, USA. dshi@cnmcresearch.org <dshi@cnmcresearch.org>
Organizational Affiliation: