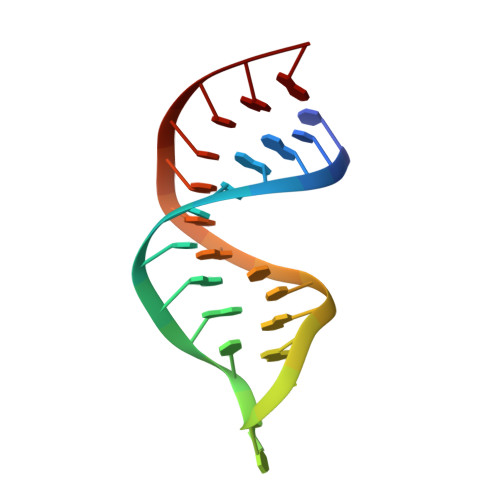
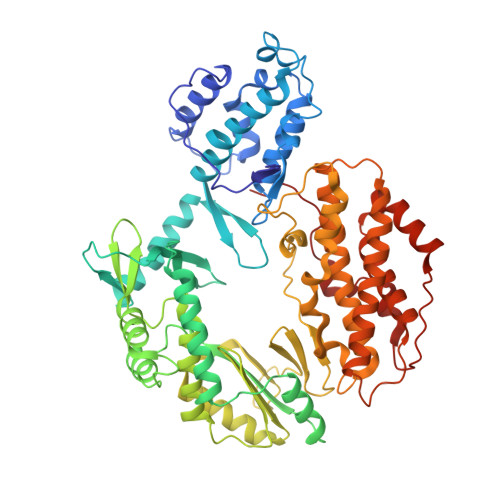
Structural basis for telomerase catalytic subunit TERT binding to RNA template and telomeric DNA.
Mitchell, M., Gillis, A., Futahashi, M., Fujiwara, H., Skordalakes, E.(2010) Nat Struct Mol Biol 17: 513-518
- PubMed: 20357774 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1777
- Primary Citation Related Structures:
3KYL - PubMed Abstract:
Telomerase is a specialized DNA polymerase that extends the 3' ends of eukaryotic linear chromosomes, a process required for genomic stability and cell viability. Here we present the crystal structure of the active Tribolium castaneum telomerase catalytic subunit, TERT, bound to an RNA-DNA hairpin designed to resemble the putative RNA-templating region and telomeric DNA. The RNA-DNA hybrid adopts a helical structure, docked in the interior cavity of the TERT ring. Contacts between the RNA template and motifs 2 and B' position the solvent-accessible RNA bases close to the enzyme active site for nucleotide binding and selectivity. Nucleic acid binding induces rigid TERT conformational changes to form a tight catalytic complex. Overall, TERT-RNA template and TERT-telomeric DNA associations are remarkably similar to those observed for retroviral reverse transcriptases, suggesting common mechanistic aspects of DNA replication between the two families of enzymes.
- Gene Expression and Regulation Program, The Wistar Institute, Philadelphia, Pennsylvania, USA.
Organizational Affiliation: