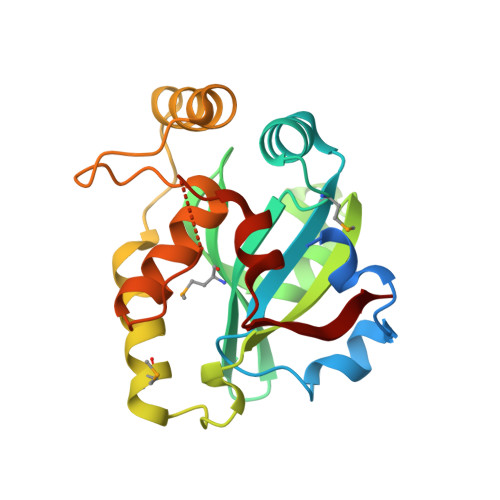
Structural Analysis of Papain-Like NlpC/P60 Superfamily Enzymes with a Circularly Permuted Topology Reveals Potential Lipid Binding Sites.
Xu, Q., Rawlings, N.D., Chiu, H.J., Jaroszewski, L., Klock, H.E., Knuth, M.W., Miller, M.D., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2011) PLoS One 6: e22013-e22013
- PubMed: 21799766 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0022013
- Primary Citation Related Structures:
3KW0 - PubMed Abstract:
NlpC/P60 superfamily papain-like enzymes play important roles in all kingdoms of life. Two members of this superfamily, LRAT-like and YaeF/YiiX-like families, were predicted to contain a catalytic domain that is circularly permuted such that the catalytic cysteine is located near the C-terminus, instead of at the N-terminus. These permuted enzymes are widespread in virus, pathogenic bacteria, and eukaryotes. We determined the crystal structure of a member of the YaeF/YiiX-like family from Bacillus cereus in complex with lysine. The structure, which adopts a ligand-induced, "closed" conformation, confirms the circular permutation of catalytic residues. A comparative analysis of other related protein structures within the NlpC/P60 superfamily is presented. Permutated NlpC/P60 enzymes contain a similar conserved core and arrangement of catalytic residues, including a Cys/His-containing triad and an additional conserved tyrosine. More surprisingly, permuted enzymes have a hydrophobic S1 binding pocket that is distinct from previously characterized enzymes in the family, indicative of novel substrate specificity. Further analysis of a structural homolog, YiiX (PDB 2if6) identified a fatty acid in the conserved hydrophobic pocket, thus providing additional insights into possible function of these novel enzymes.
- Joint Center for Structural Genomics, La Jolla, California, United States of America.
Organizational Affiliation: