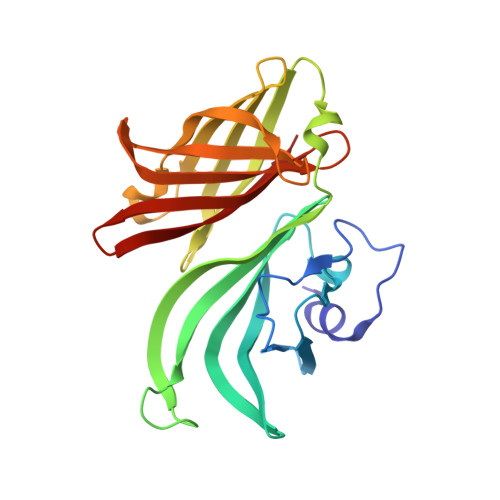
Structure of the uncomplexed Neisseria meningitidis factor H-binding protein fHbp (rLP2086).
Cendron, L., Veggi, D., Girardi, E., Zanotti, G.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 531-535
- PubMed: 21543855 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111006154
- Primary Citation Related Structures:
3KVD - PubMed Abstract:
fHbp, a highly immunogenic outer membrane protein of Neisseria meningitidis, is responsible for binding to human factor H, a multi-domain protein which is the central regulator of the alternative complement pathway. Here, the crystal structure of mature fHbp determined at 2 Å resolution is presented and is compared with the structure of the same protein in complex with factor H domains 6 and 7 recently solved using X-ray techniques. While the overall protein fold is well conserved, modifications are observed mainly in the loop regions involved in the interaction, reflecting a specific adaptation of fHbp in complexing factor H with high affinity. Such a comparison has to date been impaired by the fact that fHbp models determined by NMR show remarkable differences over the entire structure.
- Department of Biological Chemistry, University of Padua, Viale G. Colombo 3, 35121 Padua, Italy. laura.cendron@unipd.it
Organizational Affiliation: