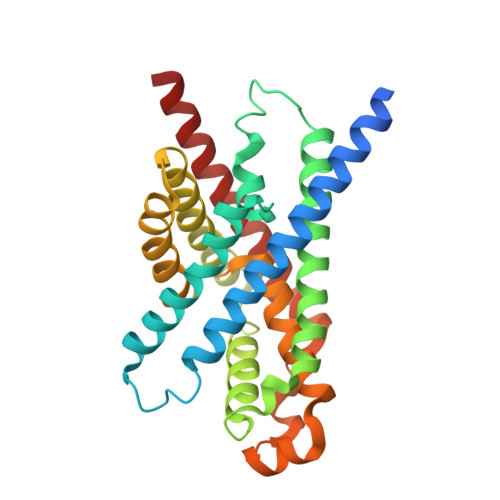
Structure and mechanism of a pentameric formate channel
Waight, A.B., Love, J., Wang, D.N.(2010) Nat Struct Mol Biol 17: 31-37
- PubMed: 20010838 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1740
- Primary Citation Related Structures:
3KLY, 3KLZ - PubMed Abstract:
Formate transport across the inner membrane is a critical step in anaerobic bacterial respiration. Members of the formate/nitrite transport protein family function to shuttle substrate across the cytoplasmic membrane. In bacterial pathogens, the nitrite transport protein is involved in protecting bacteria from peroxynitrite released by host macrophages. We have determined the 2.13-A structure of the formate channel FocA from Vibrio cholerae, which reveals a pentamer in which each monomer possesses its own substrate translocation pore. Unexpectedly, the fold of the FocA monomer resembles that found in water and glycerol channels. The selectivity filter in FocA consists of a cytoplasmic slit and a central constriction ring. A 2.5-A high-formate structure shows two formate ions bound to the cytoplasmic slit via both hydrogen bonding and van der Waals interactions, providing a structural basis for the substrate selectivity of the channel.
- The Helen L. and Martin S. Kimmel Center for Biology and Medicine, Skirball Institute of Biomolecular Medicine, New York University School of Medicine, New York, New York, USA.
Organizational Affiliation: