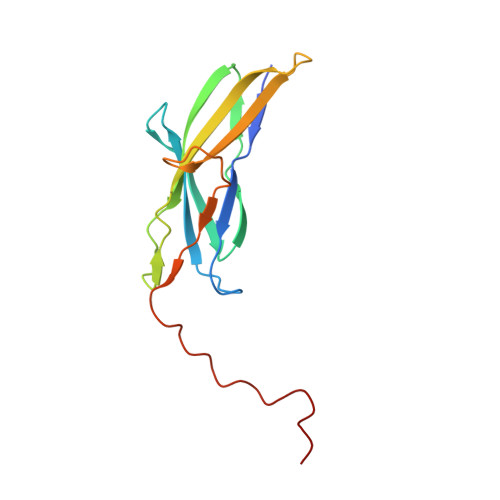
Crystal structure of the minor pilin FctB reveals determinants of Group A streptococcal pilus anchoring
Linke, C., Young, P.G., Kang, H.J., Bunker, R.D., Middleditch, M.J., Caradoc-Davies, T.T., Proft, T., Baker, E.N.(2010) J Biological Chem 285: 20381-20389
- PubMed: 20427291 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.089680
- Primary Citation Related Structures:
3KLQ - PubMed Abstract:
Cell surface pili are polymeric protein assemblies that enable bacteria to adhere to surfaces and to specific host tissues. The pili expressed by Gram-positive bacteria constitute a unique paradigm in which sortase-mediated covalent linkages join successive pilin subunits like beads on a string. These pili are formed from two or three distinct types of pilin subunit, typically encoded in small gene clusters, often with their cognate sortases. In Group A streptococci (GAS), a major pilin forms the polymeric backbone, whereas two minor pilins are located at the tip and the base. Here, we report the 1.9-A resolution crystal structure of the GAS basal pilin FctB, revealing an immunoglobulin (Ig)-like N-terminal domain with an extended proline-rich tail. Unexpected structural homology between the FctB Ig-like domain and the N-terminal domain of the GAS shaft pilin helps explain the use of the same sortase for polymerization of the shaft and its attachment to FctB. It also enabled the identification, from mass spectral data, of the lysine residue involved in the covalent linkage of FctB to the shaft. The proline-rich tail forms a polyproline-II helix that appears to be a common feature of the basal (cell wall-anchoring) pilins. Together, our results indicate distinct structural elements in the pilin proteins that play a role in selecting for the appropriate sortases and thereby help orchestrate the ordered assembly of the pilus.
- School of Biological Sciences, University of Auckland, Private Bag 92019, Auckland 1142, New Zealand.
Organizational Affiliation: