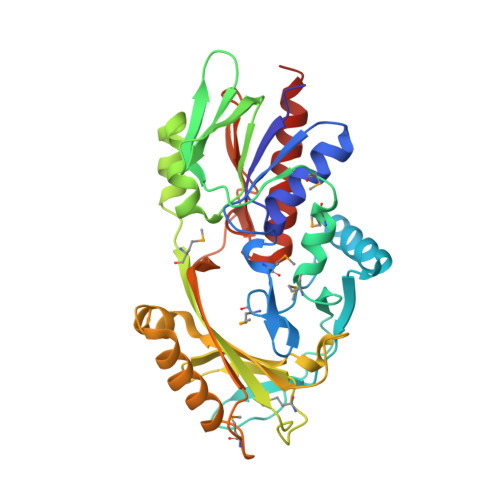
X-ray structure of P. syringae q888a4 Oxidoreductase at resolution 2.5A, Northeast Structural Genomics Consortium target PsR10
Kuzin, A.P., Chen, Y., Forouhar, F., Vorobiev, S., Acton, T., Ma, L.C., Xiao, R., Montelione, G.T., Hunt, J.F., Tong, T.To be published.