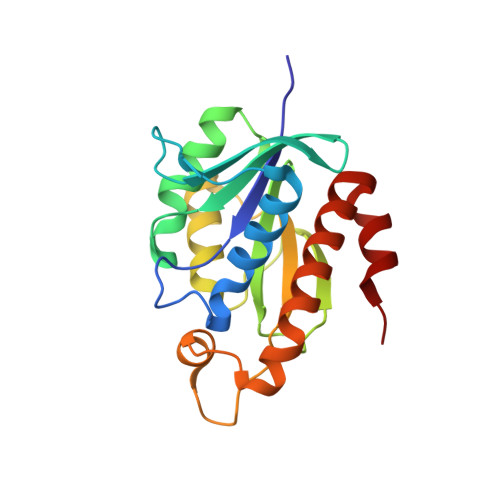
Crystal structure of peptidyl-tRNA hydrolase from mycobacterium smegmatis reveals novel features related to enzyme dynamics.
Kumar, A., Singh, N., Yadav, R., Kumar, R.P., Sharma, S., Arora, A., Singh, T.P.(2012) Int J Biochem Mol Biol 3: 58-69
- PubMed: 22509481 Search on PubMedSearch on PubMed Central
- Primary Citation Related Structures:
3KJZ - PubMed Abstract:
Peptidyl-tRNA hydrolase from Mycobacterium smegmatis is a single domain 21 kDa protein involved in the hydrolysis of prematurely produced peptidyl-tRNAs to ensure the viability of cells in bacteria, thus making it a potentially important drug target. In order to aid the development of potent drugs for controlling bacterial infections, the three-dimensional structure of peptidyl-tRNA hydrolase from Mycobacterium smegmatis has been determined. The protein adopts a compact α/β globular fold with a twisted β-sheet surrounded by α-helices. The functionally important C-terminal stretch has been unambiguously modeled for the first time in the unliganded structure of peptidyl-tRNA hydrolase. The segment, Gly138 - Val150 is mobile because it lacks significant interactions with the rest of the protein molecule. This conformational flexibility is reflected through different values of distances between a reference atom Ala147 C(α) of the segment Gly138 - Val150 to Gly114 C(α) from another segment from opposite side of the substrate binding channel in Mycobacterium smegmatis (7.8 Ǻ), Mycobacterium tuberculosis (9.5 Ǻ) and Escherichia coli (11.8 Ǻ). Similarly, the conformation of loop Gly109 - Gly117 with respect to another loop Asp95 - Asp100 also shows variability of the substrate binding cleft as the distance between Asp98 O(δ2) to Gly113 C(α) in Mycobacterium smegmatis is 4.5 Ǻ while the corresponding distances in Mycobacterium tuberculosis and Escherichia coli are 3.1 Ǻ and 6.7 Ǻ respectively. The hydrogen bonded interactions between Asn116, His22 and Asp95 indicate a stereochemically favorable arrangement of these residues for catalytic action.