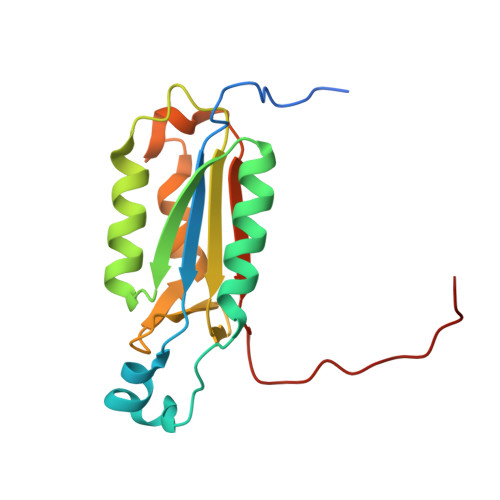
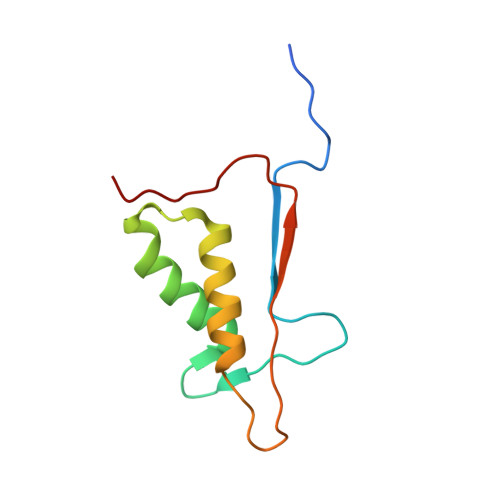
Kinetic and structural characterization of caspase-3 and caspase-8 inhibition by a novel class of irreversible inhibitors.
Wang, Z., Watt, W., Brooks, N.A., Harris, M.S., Urban, J., Boatman, D., McMillan, M., Kahn, M., Heinrikson, R.L., Finzel, B.C., Wittwer, A.J., Blinn, J., Kamtekar, S., Tomasselli, A.G.(2010) Biochim Biophys Acta 1804: 1817-1831
- PubMed: 20580860 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2010.05.007
- Primary Citation Related Structures:
3KJF, 3KJN, 3KJQ - PubMed Abstract:
Because of their central role in programmed cell death, the caspases are attractive targets for developing new therapeutics against cancer and autoimmunity, myocardial infarction and ischemic damage, and neurodegenerative diseases. We chose to target caspase-3, an executioner caspase, and caspase-8, an initiator caspase, based on the vast amount of information linking their functions to diseases. Through a structure-based drug design approach, a number of novel beta-strand peptidomimetic compounds were synthesized. Kinetic studies of caspase-3 and caspase-8 inhibition were carried out with these urazole ring-containing irreversible peptidomimetics and a known irreversible caspase inhibitor, Z-VAD-fmk. Using a stopped-flow fluorescence assay, we were able to determine individual kinetic parameters of caspase-3 and caspase-8 inhibition by these inhibitors. Z-VAD-fmk and the peptidomimetic inhibitors inhibit caspase-3 and caspase-8 via a three-step kinetic mechanism. Inhibition of both caspase-3 and caspase-8 by Z-VAD-fmk and of caspase-3 by the peptidomimetic inhibitors proceeds via two rapid equilibrium steps followed by a relatively fast inactivation step. However, caspase-8 inhibition by the peptidomimetics goes through a rapid equilibrium step, a slow-binding reversible step, and an extremely slow inactivation step. The crystal structures of inhibitor complexes of caspases-3 and -8 validate the design of the inhibitors by illustrating in detail how they mimic peptide substrates. One of the caspase-8 structures also shows binding at a secondary, allosteric site, providing a possible route to the development of noncovalent small molecule modulators of caspase activity.
- Oligonucleotide Therapeutics Unit, Pfizer, Inc., 620 Memorial Drive, Cambridge, MA 02139, USA. zhigang.wang@pfizer.com
Organizational Affiliation: