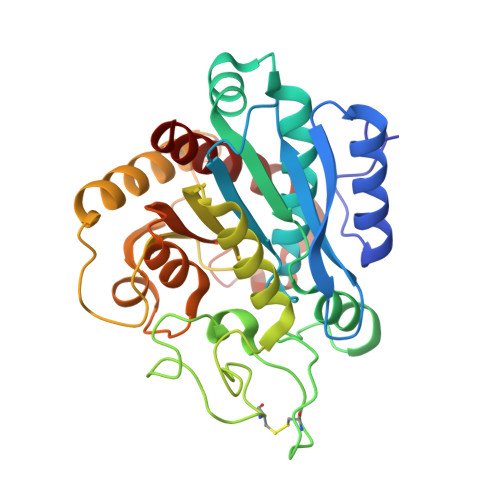
Structural and Functional Analysis of the Complex between Citrate and the Zinc Peptidase Carboxypeptidase A
Fernandez, D., Boix, E., Pallares, I., Aviles, F.X., Vendrell, J.(2011) Enzyme Res 2011: 128676-128676
- PubMed: 21804935 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4061/2011/128676
- Primary Citation Related Structures:
3KGQ - PubMed Abstract:
A high-resolution carboxypeptidase-Zn(2+)-citrate complex was studied by X-ray diffraction and enzyme kinetics for the first time. The citrate molecule acts as a competitive inhibitor of this benchmark zinc-dependent peptidase, chelating the catalytic zinc ion in the active site of the enzyme and inducing a conformational change such that carboxypeptidase adopts the conformation expected to occur by substrate binding. Citrate adopts an extended conformation with half of the molecule facing the zinc ion, while the other half is docked in the S1' hydrophobic specificity pocket of the enzyme, in contrast with the binding mode expected for a substrate like phenylalanine or a peptidomimetic inhibitor like benzylsuccinic acid. Combined structural and enzymatic analysis describes the characteristics of the binding of this ligand that, acting against physiologically relevant zinc-dependent proteases, may serve as a general model in the design of new drug-protecting molecules for the oral delivery of drugs of peptide origin.
- Departament de Bioquímica i Biologia Molecular, Facultat de Biociències, Universitat Autònoma de Barcelona, 08193 Bellaterra, Spain.
Organizational Affiliation: